Week 1, Session 3 — Data types, tidy data, accuracy and precision
Course 1 — #courses
Note
Workflow labs use the variant template: Goal → Approach → Execution → Check → Report.
Learning objectives
- Classify a variable as nominal, ordinal, interval, or ratio and explain why the distinction dictates what summaries are legal.
- Reshape a rectangular dataset between wide and long form with
pivot_longer()andpivot_wider(). - Separate accuracy, precision, and bias by simulation, and say why a precise instrument can still be wrong.
Prerequisites
Lab 1.2 (toolchain).
Background
Statistics is largely the study of variation, and variation must be measured. Before we calculate anything we need to say what kind of variable we are looking at. A nominal variable labels groups without order. An ordinal variable orders them but does not quantify the distance between them. Interval and ratio variables do quantify distances, and ratio variables have a true zero. Tools that average an ordinal variable or compute a standard deviation on a category code produce numbers, but not answers.
Tidy data is a convention, not a requirement, but adopting it pays immediate dividends: every variable gets a column, every observation a row, and every value a cell. Real datasets arrive in other shapes, so a large part of the job is reshaping them until they conform. The tools are pivot_longer() and pivot_wider(), and they are the two functions you will use most often in a working week.
The accuracy/precision/bias triangle is borrowed from measurement theory. Accuracy is how close a measurement is to truth on average. Precision is how tightly repeated measurements cluster. Bias is a systematic offset from truth. An instrument can be precise but biased (tight cluster off-target), accurate but imprecise (wide cluster centred on truth), or — the goal — both.
Setup
1. Goal
Move a small simulated clinical dataset between wide and long forms and quantify accuracy, precision, and bias in a simulated measurement process.
2. Approach
We simulate three repeat measurements of blood pressure on twelve patients at two visits. This is a realistic wide-format dataset with a within-patient structure.
clinic <- tibble(
id = sprintf("P%02d", 1:12),
sex = sample(c("F", "M"), 12, replace = TRUE),
visit1_sbp1 = rnorm(12, 140, 10),
visit1_sbp2 = rnorm(12, 140, 10),
visit1_sbp3 = rnorm(12, 140, 10),
visit2_sbp1 = rnorm(12, 135, 10),
visit2_sbp2 = rnorm(12, 135, 10),
visit2_sbp3 = rnorm(12, 135, 10)
)
clinic |> select(id, starts_with("visit1")) |> head(3)# A tibble: 3 × 4
id visit1_sbp1 visit1_sbp2 visit1_sbp3
<chr> <dbl> <dbl> <dbl>
1 P01 155. 116. 145.
2 P02 139. 153. 147.
3 P03 160. 137. 150.3. Execution
Reshape to a long form where each row is one reading.
# A tibble: 6 × 5
id sex visit reading sbp
<chr> <chr> <fct> <int> <dbl>
1 P01 F V1 1 155.
2 P01 F V1 2 116.
3 P01 F V1 3 145.
4 P01 F V2 1 143.
5 P01 F V2 2 136.
6 P01 F V2 3 138.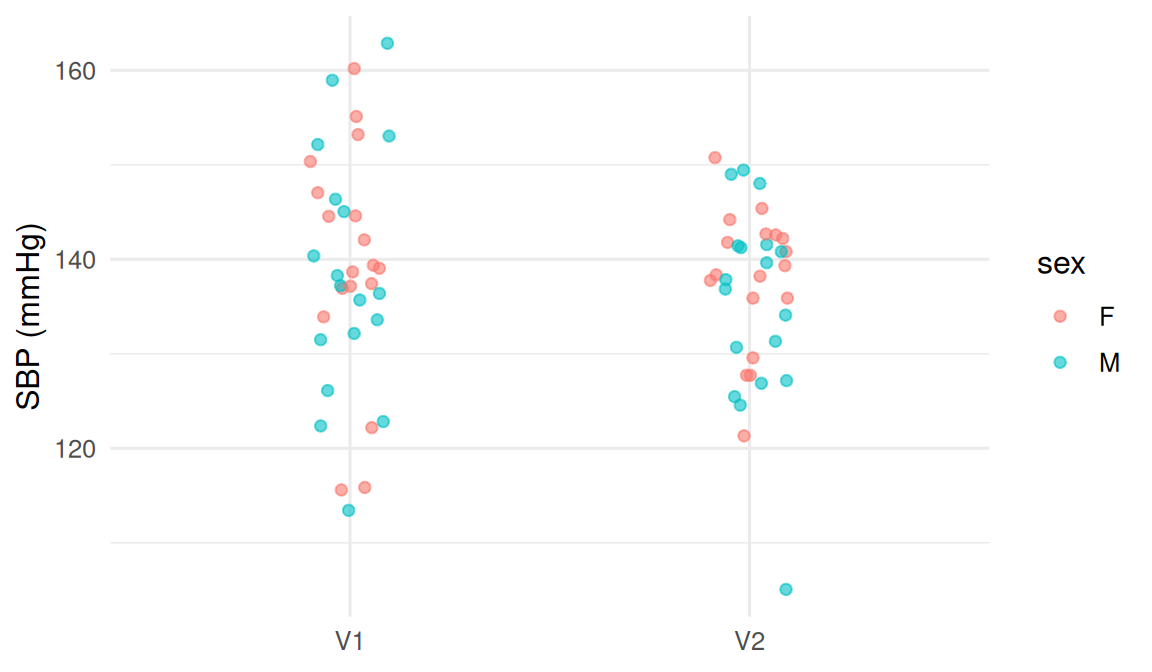
The long format makes it trivial to summarise by visit, by patient, or by visit-and-patient. The same plot in wide format would require custom code for each visit column.
# A tibble: 6 × 4
id sex V1 V2
<chr> <chr> <dbl> <dbl>
1 P01 F 138. 139.
2 P02 F 146. 137.
3 P03 F 149. 136.
4 P04 F 132. 139.
5 P05 M 145. 119.
6 P06 M 146. 140.4. Check
Simulate an instrument with a known truth (say, a calibrated reference of 120) and compare three candidate devices with different accuracy/precision profiles.
truth <- 120
n <- 500
devices <- tibble(
device_A = rnorm(n, mean = 120, sd = 1), # accurate & precise
device_B = rnorm(n, mean = 125, sd = 1), # precise, biased
device_C = rnorm(n, mean = 120, sd = 5) # accurate on average, imprecise
) |>
pivot_longer(everything(), names_to = "device", values_to = "reading")
summary_table <- devices |>
group_by(device) |>
summarise(mean = mean(reading),
sd = sd(reading),
bias = mean(reading) - truth,
rmse = sqrt(mean((reading - truth)^2)))
summary_table# A tibble: 3 × 5
device mean sd bias rmse
<chr> <dbl> <dbl> <dbl> <dbl>
1 device_A 120. 0.974 -0.0280 0.974
2 device_B 125. 1.02 4.95 5.06
3 device_C 120. 4.83 -0.0819 4.83 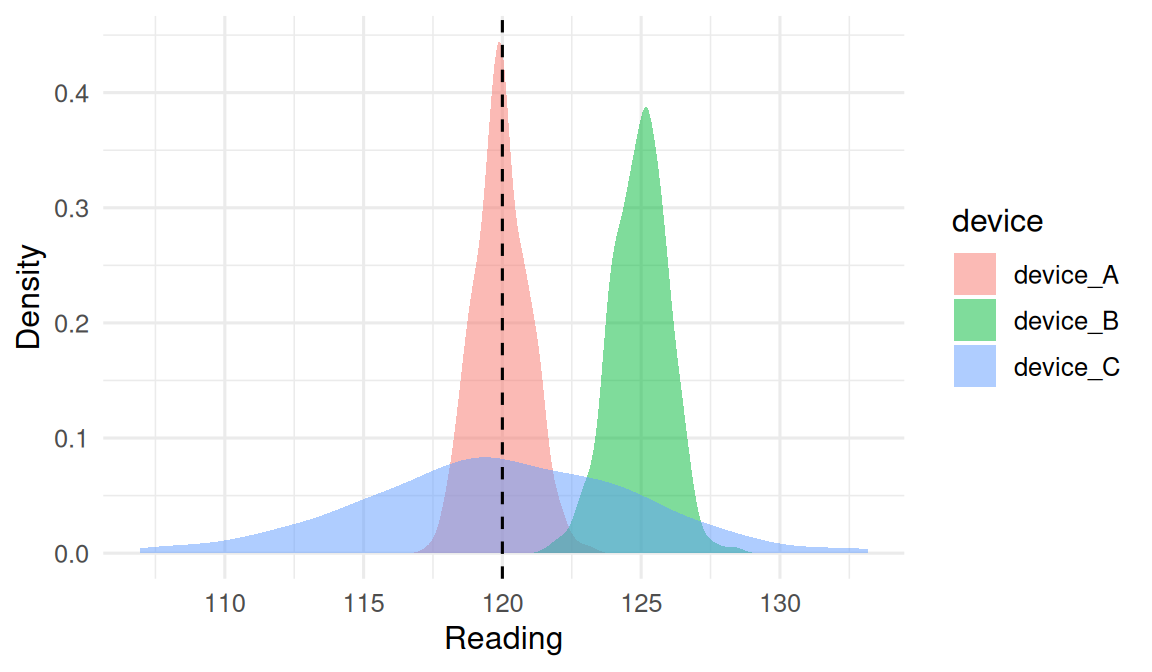
5. Report
In a simulated calibration experiment (n = 500 draws per device), the unbiased precise device had bias = -0.03 mmHg and SD = 0.97; the biased precise device had bias = 4.95 and the same SD; the imprecise unbiased device had bias = -0.08 but SD = 4.83. Root mean squared error captures accuracy overall and is the quantity to minimise.
The lesson is that bias and variance are separately identifiable. A device that is consistent is not necessarily correct, and a device that is correct on average may still be unusable on a single reading.
Common pitfalls
- Treating ordinal codes (1 = mild, 2 = moderate, 3 = severe) as if the steps were equal.
- Forcing data to stay wide because that is how the spreadsheet happened to arrive.
- Judging a measurement by precision alone. A small SD with a big bias is still the wrong number.
- Using the mean to summarise a nominal variable.
Further reading
- Wickham H (2014), Tidy Data. Journal of Statistical Software.
- Altman DG, Bland JM (1983), Measurement in medicine.
Session info
R version 4.4.1 (2024-06-14)
Platform: x86_64-pc-linux-gnu
Running under: Ubuntu 24.04.4 LTS
Matrix products: default
BLAS: /usr/lib/x86_64-linux-gnu/openblas-pthread/libblas.so.3
LAPACK: /usr/lib/x86_64-linux-gnu/openblas-pthread/libopenblasp-r0.3.26.so; LAPACK version 3.12.0
locale:
[1] LC_CTYPE=C.UTF-8 LC_NUMERIC=C LC_TIME=C.UTF-8
[4] LC_COLLATE=C.UTF-8 LC_MONETARY=C.UTF-8 LC_MESSAGES=C.UTF-8
[7] LC_PAPER=C.UTF-8 LC_NAME=C LC_ADDRESS=C
[10] LC_TELEPHONE=C LC_MEASUREMENT=C.UTF-8 LC_IDENTIFICATION=C
time zone: UTC
tzcode source: system (glibc)
attached base packages:
[1] stats graphics grDevices utils datasets methods base
other attached packages:
[1] lubridate_1.9.5 forcats_1.0.1 stringr_1.6.0 dplyr_1.2.1
[5] purrr_1.2.2 readr_2.2.0 tidyr_1.3.2 tibble_3.3.1
[9] ggplot2_4.0.3 tidyverse_2.0.0
loaded via a namespace (and not attached):
[1] gtable_0.3.6 jsonlite_2.0.0 compiler_4.4.1 tidyselect_1.2.1
[5] scales_1.4.0 yaml_2.3.12 fastmap_1.2.0 R6_2.6.1
[9] labeling_0.4.3 generics_0.1.4 knitr_1.51 htmlwidgets_1.6.4
[13] pillar_1.11.1 RColorBrewer_1.1-3 tzdb_0.5.0 rlang_1.2.0
[17] utf8_1.2.6 stringi_1.8.7 xfun_0.57 S7_0.2.2
[21] otel_0.2.0 timechange_0.4.0 cli_3.6.6 withr_3.0.2
[25] magrittr_2.0.5 digest_0.6.39 grid_4.4.1 hms_1.1.4
[29] lifecycle_1.0.5 vctrs_0.7.3 evaluate_1.0.5 glue_1.8.1
[33] farver_2.1.2 rmarkdown_2.31 tools_4.4.1 pkgconfig_2.0.3
[37] htmltools_0.5.9