Week 3, Session 4 — One-sample t-test and one-proportion test
Course 1 — #courses
Note
Inference labs use the five-step template: Hypothesis → Visualise → Assumptions → Conduct → Conclude.
Learning objectives
- State a one-sample null and alternative hypothesis for a continuous outcome and for a proportion.
- Run
t.test(mu = )andprop.test()/binom.test()and interpret the output. - Apply the full five-step inference template in one working pass.
Prerequisites
Labs 2.5, 3.3.
Background
The one-sample t-test asks whether the mean of a continuous outcome differs from a pre-specified value μ₀. It is the inference analogue of Table 1: a single-column calculation asking whether the sample is consistent with a named population mean. The test statistic is (x̄ − μ₀) / (s/√n), compared to a t distribution on n − 1 degrees of freedom.
The one-proportion test does the same for a binary outcome. When the sample is large we use prop.test(), which is a Wald-type test based on the normal approximation to the binomial; when the sample is small, binom.test() gives an exact test based on the binomial PMF. Both return a p-value, a point estimate, and a CI.
These are the simplest tests in the course, but the five-step template — hypothesis, visualise, assumptions, conduct, conclude — is the same template used in every later inference lab. Getting the template into muscle memory here pays dividends throughout the rest of the course.
Setup
1. Hypothesis
Scenario A (continuous): fasting blood glucose in a random sample of n = 40 patients from a clinic. Reference mean μ₀ = 95 mg/dL in a healthy population. Two-sided test.
- H0: μ = 95
- H1: μ ≠ 95
- α = 0.05
Scenario B (proportion): of 120 patients offered a new screening programme, k = 78 accepted. Reference acceptance rate π₀ = 0.60.
- H0: π = 0.60
- H1: π ≠ 0.60
- α = 0.05
2. Visualise
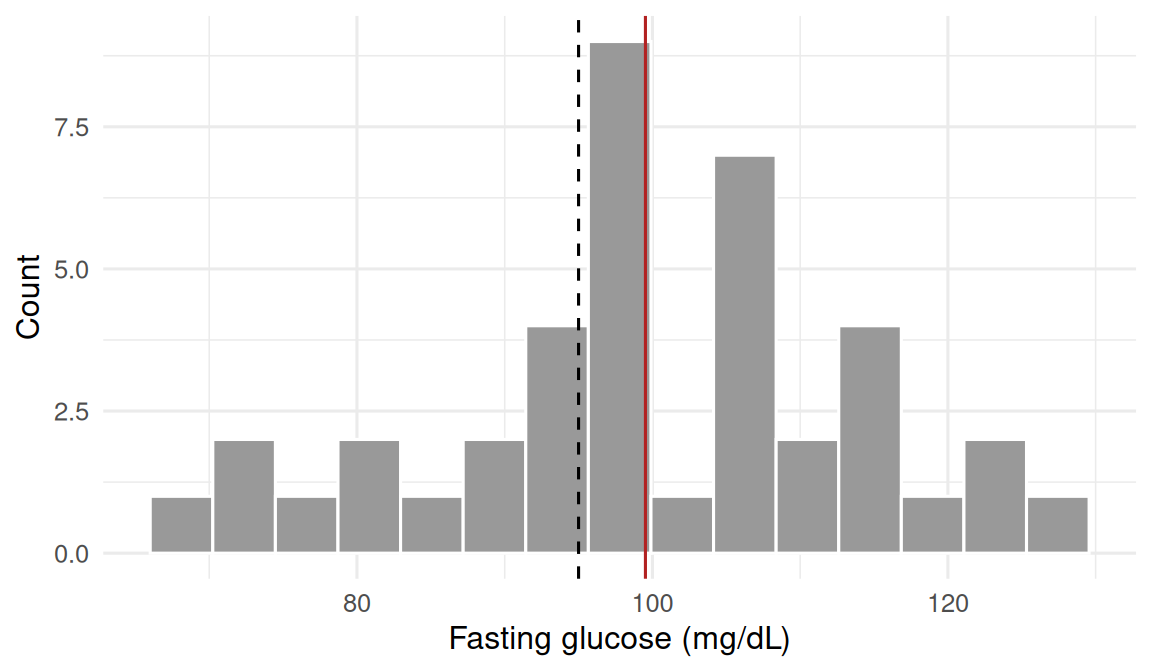
3. Assumptions
t-test: approximate normality of the sample mean, which — by the central limit theorem — holds for n = 40 unless the underlying distribution is strongly skewed. Check with a Q-Q plot.
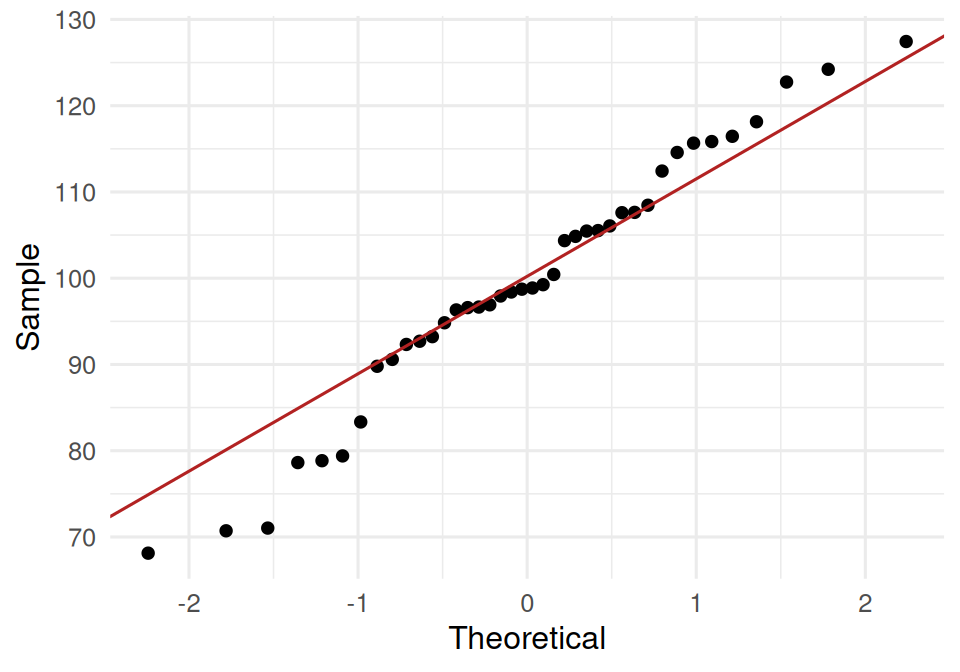
Proportion test: independent binary trials, same success probability. For prop.test the large-sample normal approximation needs np₀ ≥ 5 and n(1 − p₀) ≥ 5; both are satisfied.
4. Conduct
One Sample t-test
data: glucose
t = 1.9513, df = 39, p-value = 0.05824
alternative hypothesis: true mean is not equal to 95
95 percent confidence interval:
94.83431 104.21683
sample estimates:
mean of x
99.52557 $prop_test
1-sample proportions test with continuity correction
data: accept out of total, null probability 0.6
X-squared = 1.0503, df = 1, p-value = 0.3054
alternative hypothesis: true p is not equal to 0.6
95 percent confidence interval:
0.5569559 0.7332917
sample estimates:
p
0.65
$binom_test
Exact binomial test
data: accept and total
number of successes = 78, number of trials = 120, p-value = 0.3054
alternative hypothesis: true probability of success is not equal to 0.6
95 percent confidence interval:
0.5576053 0.7347977
sample estimates:
probability of success
0.65 5. Concluding statement
Scenario A. In a sample of 40 patients, mean fasting glucose was 99.5 mg/dL (SD 14.7). The one-sample t-test against μ₀ = 95 gave t(39) = 1.95, p = 0.058, mean difference 4.5 (95% CI -0.2 to 9.2).
Scenario B. Acceptance was 78 of 120 (65%); the exact one-proportion test against π₀ = 0.60 gave p = 0.31, 95% CI 0.56 to 0.73.
The two scenarios share a template. The glossary changes — mean and SD for continuous, count and proportion for binary — but the five steps are the same.
Common pitfalls
- Running a one-sided test to “rescue” a borderline two-sided p-value.
- Reporting the p-value without the mean difference and its CI.
- Using
prop.testwhen the expected count of either outcome is very small; usebinom.testthere.
Further reading
- Rosner B. Fundamentals of Biostatistics, chapter on one-sample inference.
- Altman DG, Confidence intervals rather than p values, BMJ 1986.
Session info
R version 4.4.1 (2024-06-14)
Platform: x86_64-pc-linux-gnu
Running under: Ubuntu 24.04.4 LTS
Matrix products: default
BLAS: /usr/lib/x86_64-linux-gnu/openblas-pthread/libblas.so.3
LAPACK: /usr/lib/x86_64-linux-gnu/openblas-pthread/libopenblasp-r0.3.26.so; LAPACK version 3.12.0
locale:
[1] LC_CTYPE=C.UTF-8 LC_NUMERIC=C LC_TIME=C.UTF-8
[4] LC_COLLATE=C.UTF-8 LC_MONETARY=C.UTF-8 LC_MESSAGES=C.UTF-8
[7] LC_PAPER=C.UTF-8 LC_NAME=C LC_ADDRESS=C
[10] LC_TELEPHONE=C LC_MEASUREMENT=C.UTF-8 LC_IDENTIFICATION=C
time zone: UTC
tzcode source: system (glibc)
attached base packages:
[1] stats graphics grDevices utils datasets methods base
other attached packages:
[1] lubridate_1.9.5 forcats_1.0.1 stringr_1.6.0 dplyr_1.2.1
[5] purrr_1.2.2 readr_2.2.0 tidyr_1.3.2 tibble_3.3.1
[9] ggplot2_4.0.3 tidyverse_2.0.0
loaded via a namespace (and not attached):
[1] gtable_0.3.6 jsonlite_2.0.0 compiler_4.4.1 tidyselect_1.2.1
[5] scales_1.4.0 yaml_2.3.12 fastmap_1.2.0 R6_2.6.1
[9] labeling_0.4.3 generics_0.1.4 knitr_1.51 htmlwidgets_1.6.4
[13] pillar_1.11.1 RColorBrewer_1.1-3 tzdb_0.5.0 rlang_1.2.0
[17] utf8_1.2.6 stringi_1.8.7 xfun_0.57 S7_0.2.2
[21] otel_0.2.0 timechange_0.4.0 cli_3.6.6 withr_3.0.2
[25] magrittr_2.0.5 digest_0.6.39 grid_4.4.1 hms_1.1.4
[29] lifecycle_1.0.5 vctrs_0.7.3 evaluate_1.0.5 glue_1.8.1
[33] farver_2.1.2 rmarkdown_2.31 tools_4.4.1 pkgconfig_2.0.3
[37] htmltools_0.5.9