Week 4, Session 3 — Pearson and Spearman correlation
Course 1 — #courses
Note
Inference labs use the five-step template: Hypothesis → Visualise → Assumptions → Conduct → Conclude.
Learning objectives
- Compute Pearson and Spearman correlations with
cor.test()and interpret the coefficient and its CI. - Distinguish linear association from monotonic association.
- Use visualisation to diagnose situations where a correlation coefficient misleads (Anscombe, DatasauRus).
Prerequisites
Week 1, Lab 2.5.
Background
Pearson’s r measures linear association. It is the standardised covariance of two variables; it is sensitive to outliers and assumes a roughly linear, bivariate-normal relationship. Spearman’s ρ measures monotonic association. It is Pearson’s r applied to the ranks of the variables; it is robust to outliers and to non-linear monotonic transformations.
Neither coefficient captures non-linear, non-monotonic relationships well. Anscombe’s quartet and the DatasauRus dozen are the canonical demonstrations that a coefficient without a plot can lie — datasets engineered to have identical Pearson r but radically different shapes. The conclusion is not to abandon correlations but never to report one without first plotting the data.
A correlation coefficient is not a causal quantity. Two variables can be perfectly correlated without one causing the other — a lurking variable, a common cause, a coincidence of sample selection. The word “correlation” is often used loosely in the press to mean “weak causation”; in this course it means exactly what cor.test() returns.
Setup
1. Hypothesis
Two scenarios:
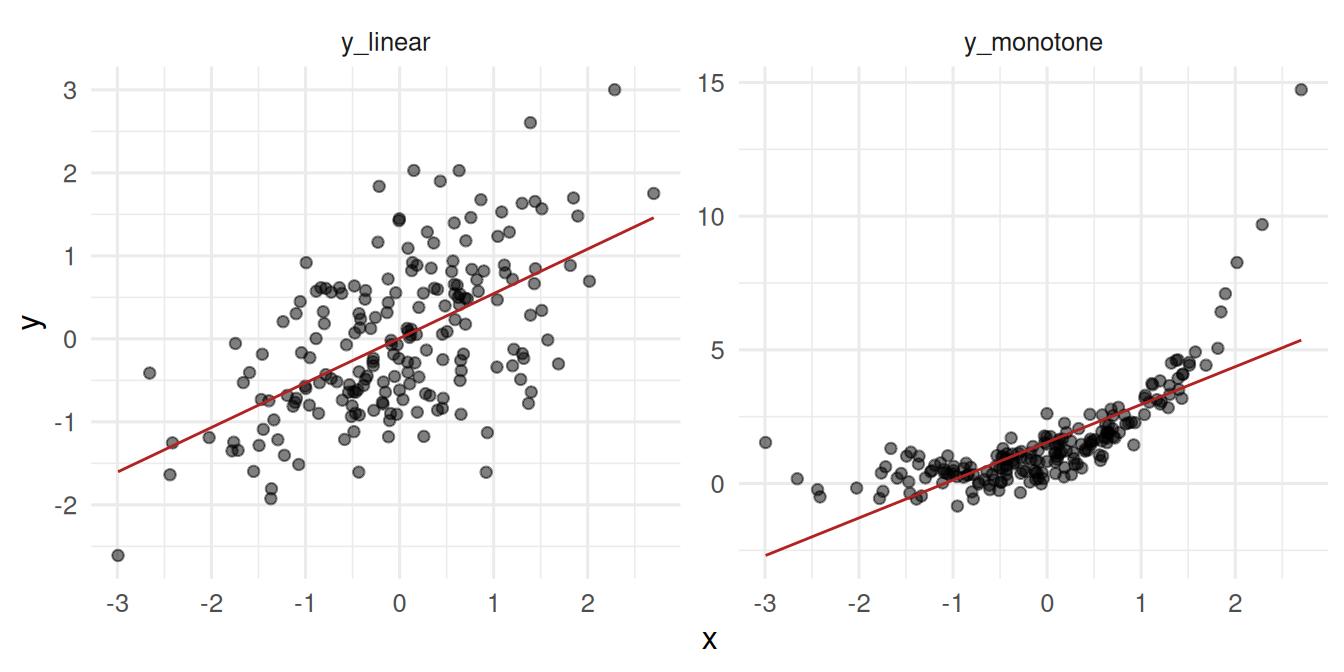
- A linear relationship between two simulated variables.
- A monotonic but non-linear relationship where Pearson misleads.
H0 in each case: ρ = 0. H1: ρ ≠ 0.
2. Visualise
3. Assumptions
Pearson assumes linearity and roughly bivariate-normal data. Spearman assumes monotonicity. Neither handles non-monotonic relationships. For both, the observations must be independent; both are sensitive to extreme sample-size-induced significance when n is huge.
4. Conduct
p1_pearson <- cor.test(x, y_linear, method = "pearson")
p1_spearman <- cor.test(x, y_linear, method = "spearman", exact = FALSE)
p2_pearson <- cor.test(x, y_monotone, method = "pearson")
p2_spearman <- cor.test(x, y_monotone, method = "spearman", exact = FALSE)
tibble(
relationship = c("linear", "linear", "monotone", "monotone"),
method = c("Pearson", "Spearman", "Pearson", "Spearman"),
r_or_rho = c(p1_pearson$estimate, p1_spearman$estimate,
p2_pearson$estimate, p2_spearman$estimate),
p = c(p1_pearson$p.value, p1_spearman$p.value,
p2_pearson$p.value, p2_spearman$p.value)
)# A tibble: 4 × 4
relationship method r_or_rho p
<chr> <chr> <dbl> <dbl>
1 linear Pearson 0.570 1.27e-18
2 linear Spearman 0.528 8.89e-16
3 monotone Pearson 0.762 2.90e-39
4 monotone Spearman 0.833 7.32e-53The monotonic non-linear relationship gives a much higher Spearman than Pearson — the rank correlation sees the perfect monotonicity.
The DatasauRus demonstration
# A tibble: 6 × 6
dataset mean_x mean_y sd_x sd_y r
<chr> <dbl> <dbl> <dbl> <dbl> <dbl>
1 away 54.3 47.8 16.8 26.9 -0.0641
2 bullseye 54.3 47.8 16.8 26.9 -0.0686
3 circle 54.3 47.8 16.8 26.9 -0.0683
4 dino 54.3 47.8 16.8 26.9 -0.0645
5 dots 54.3 47.8 16.8 26.9 -0.0603
6 h_lines 54.3 47.8 16.8 26.9 -0.0617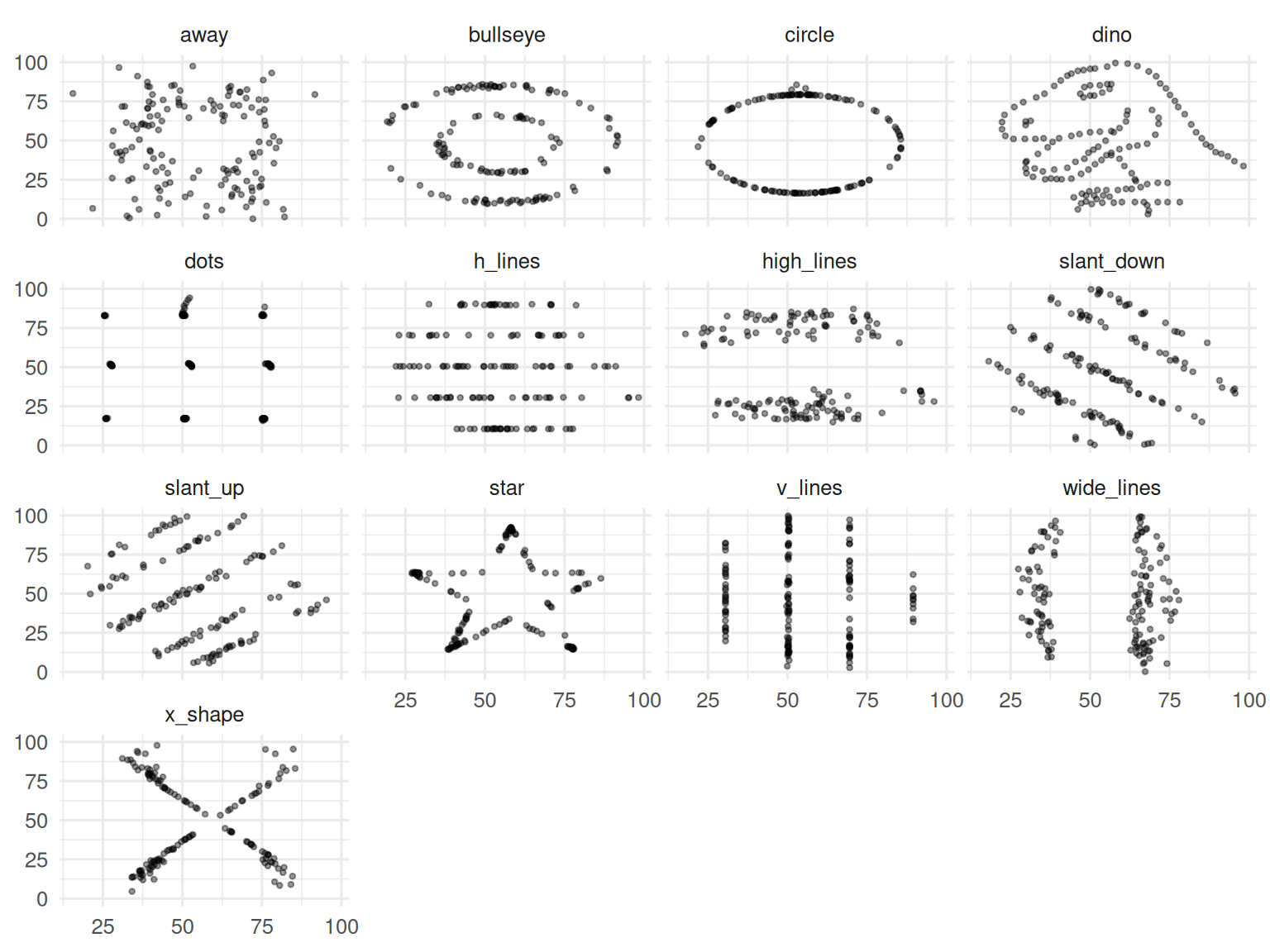
Thirteen datasets; all have mean x ≈ 54, mean y ≈ 47, SDs ≈ 17 and 26, and Pearson r ≈ −0.06. The shapes are wildly different.
5. Concluding statement
In the linear case, Pearson r = 0.57 (p = 1.3^{-18}) and Spearman ρ = 0.53; the two agreed. In the monotonic non-linear case, Pearson r = 0.76 was substantially smaller than Spearman ρ = 0.83, revealing that the association was strong but not linear. The DatasauRus dozen reproduced identical summary statistics across 13 radically different shapes, reinforcing the rule: never report a correlation without the scatter plot.
Correlation is a single number. Its honest use requires a picture alongside.
Common pitfalls
- Reporting a Pearson r on data with an obvious curvilinear shape.
- Using r to compare groups with different sample sizes; small r can be “significant” at huge n.
- Inferring causation from a correlation.
- Computing correlations on data that include outliers without a robustness check.
Further reading
- Anscombe FJ (1973). Graphs in Statistical Analysis.
- Matejka J, Fitzmaurice G (2017). Same stats, different graphs: generating datasets with varied appearance and identical statistics.
Session info
R version 4.4.1 (2024-06-14)
Platform: x86_64-pc-linux-gnu
Running under: Ubuntu 24.04.4 LTS
Matrix products: default
BLAS: /usr/lib/x86_64-linux-gnu/openblas-pthread/libblas.so.3
LAPACK: /usr/lib/x86_64-linux-gnu/openblas-pthread/libopenblasp-r0.3.26.so; LAPACK version 3.12.0
locale:
[1] LC_CTYPE=C.UTF-8 LC_NUMERIC=C LC_TIME=C.UTF-8
[4] LC_COLLATE=C.UTF-8 LC_MONETARY=C.UTF-8 LC_MESSAGES=C.UTF-8
[7] LC_PAPER=C.UTF-8 LC_NAME=C LC_ADDRESS=C
[10] LC_TELEPHONE=C LC_MEASUREMENT=C.UTF-8 LC_IDENTIFICATION=C
time zone: UTC
tzcode source: system (glibc)
attached base packages:
[1] stats graphics grDevices utils datasets methods base
other attached packages:
[1] datasauRus_0.1.9 lubridate_1.9.5 forcats_1.0.1 stringr_1.6.0
[5] dplyr_1.2.1 purrr_1.2.2 readr_2.2.0 tidyr_1.3.2
[9] tibble_3.3.1 ggplot2_4.0.3 tidyverse_2.0.0
loaded via a namespace (and not attached):
[1] Matrix_1.7-0 gtable_0.3.6 jsonlite_2.0.0 compiler_4.4.1
[5] tidyselect_1.2.1 splines_4.4.1 scales_1.4.0 yaml_2.3.12
[9] fastmap_1.2.0 lattice_0.22-6 R6_2.6.1 labeling_0.4.3
[13] generics_0.1.4 knitr_1.51 htmlwidgets_1.6.4 pillar_1.11.1
[17] RColorBrewer_1.1-3 tzdb_0.5.0 rlang_1.2.0 utf8_1.2.6
[21] stringi_1.8.7 xfun_0.57 S7_0.2.2 otel_0.2.0
[25] timechange_0.4.0 cli_3.6.6 mgcv_1.9-1 withr_3.0.2
[29] magrittr_2.0.5 digest_0.6.39 grid_4.4.1 hms_1.1.4
[33] nlme_3.1-164 lifecycle_1.0.5 vctrs_0.7.3 evaluate_1.0.5
[37] glue_1.8.1 farver_2.1.2 rmarkdown_2.31 tools_4.4.1
[41] pkgconfig_2.0.3 htmltools_0.5.9