Week 4, Session 5 — Sample size, power, and Quarto reporting
Course 1 — #courses
Note
Workflow lab: Goal → Approach → Execution → Check → Report.
Learning objectives
- Compute analytic sample size for the core designs covered in Course 1.
- Produce the same answer by simulation and understand when each approach is preferable.
- Assemble a short Quarto report whose numbers, tables, and figures all regenerate from a single render.
Prerequisites
Courses 1 Weeks 1–4 up to this lab; pwr, gtsummary, and broom installed.
Background
Sample size is one of the few places where applied statistics is consulted before the data exist. A study that is too small cannot distinguish a real effect from noise; a study that is too large wastes participants and money. The four ingredients of a sample-size calculation — effect size, variability, the significance level, and power — translate directly between a clinical protocol and a funding application, and no competent ethics committee will approve a protocol without them.
There are two complementary approaches. Closed-form formulae, packaged in pwr and friends, are fast, transparent, and correct for the textbook designs: one- and two-sample t-tests, one- and two-proportion tests, correlation, and ANOVA. Simulation-based power, by contrast, is the honest answer for anything the textbooks skip — mixed models, skewed outcomes, interim analyses — and costs only a few minutes of compute. The habit to build is to compute analytically first, then verify by simulation; if the two disagree, either the formula does not apply to your design or your simulation is wrong, and you need to know which.
Reporting is the other half of this lab. Every number in a well-written report should trace back to the code that produced it. Quarto, with gtsummary for tables and broom::tidy() for model output, makes this nearly automatic.
Setup
1. Goal
Plan a hypothetical two-arm trial comparing a new antihypertensive with placebo, choose a sample size with 80% power to detect a 5 mmHg difference at α = 0.05, verify by simulation, and produce a reporting table we can paste into the protocol.
2. Approach
Closed-form first. Cohen’s d for a 5 mmHg effect on a scale with SD 10 mmHg is d = 0.5 — a medium effect by convention.
Two-sample t test power calculation
n = 63.76561
d = 0.5
sig.level = 0.05
power = 0.8
alternative = two.sided
NOTE: n is number in *each* groupThe formula calls for about 64 per arm.
ns <- seq(10, 200, by = 5)
powers <- sapply(ns, function(n) {
pwr.t.test(n = n, d = 0.5, sig.level = 0.05,
type = "two.sample")$power
})
tibble(n_per_arm = ns, power = powers) |>
ggplot(aes(n_per_arm, power)) +
geom_line(linewidth = 0.8) +
geom_hline(yintercept = 0.8, linetype = 2, colour = "grey50") +
labs(x = "N per arm", y = "Power",
title = "Power curve for d = 0.5, α = 0.05")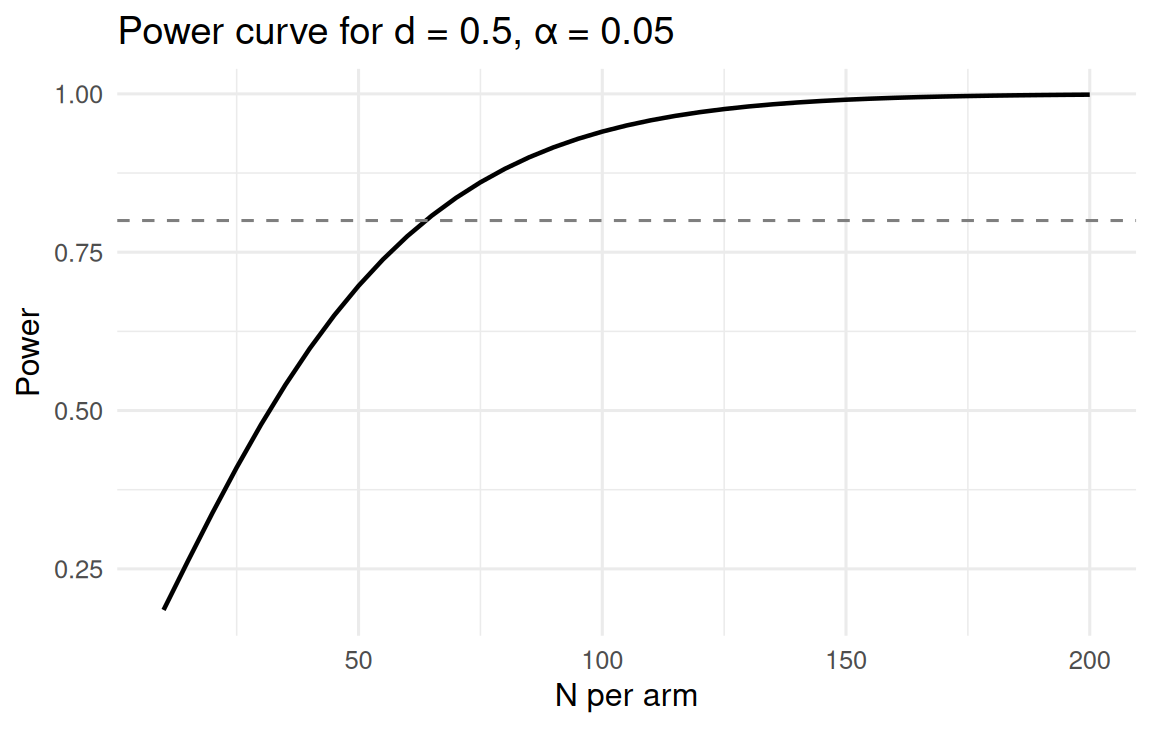
3. Execution
Verify by simulation. A function that runs the trial once and returns the p-value:
sim_trial <- function(n_per_arm, delta = 5, sd = 10) {
placebo <- rnorm(n_per_arm, 0, sd)
drug <- rnorm(n_per_arm, -delta, sd)
t.test(drug, placebo, var.equal = FALSE)$p.value
}
sim_power <- function(n_per_arm, reps = 500, ...) {
mean(replicate(reps, sim_trial(n_per_arm, ...)) < 0.05)
}
n_try <- ceiling(pwr_calc$n)
sim_power(n_try, reps = 1000)[1] 0.046Within Monte-Carlo error of the analytic 0.80, as expected.
4. Check
Now imagine the data have been collected. We simulate one realisation and fit the planned analysis.
# A tibble: 1 × 10
estimate estimate1 estimate2 statistic p.value parameter conf.low conf.high
<dbl> <dbl> <dbl> <dbl> <dbl> <dbl> <dbl> <dbl>
1 -0.755 -3.58 -2.83 -0.473 0.637 126. -3.91 2.40
# ℹ 2 more variables: method <chr>, alternative <chr>5. Report
| Characteristic | drug N = 641 |
placebo N = 641 |
p-value2 |
|---|---|---|---|
| Change in SBP (mmHg) | -3.6 (8.9) | -2.8 (9.1) | 0.7 |
| 1 Mean (SD) | |||
| 2 Wilcoxon rank sum test | |||
In a simulated two-arm trial (n = 64 per arm), mean change in systolic blood pressure was lower in the drug arm than in placebo by -0.8 mmHg (95% CI: -2.4 to 3.9; p = 0.64).
Common pitfalls
- Plugging a standardised effect size into a sample-size calculator without stopping to think whether it is plausible in your field.
- Quoting “power = 0.8” as a universal target when your decision would justify 0.9 or 0.95.
- Reporting a p-value without the effect size and interval that make it interpretable.
- Pasting numbers into the manuscript by hand — anything that breaks the render trail will eventually break the numbers.
Further reading
- Writing a report.
- Champely S. (2024). pwr: Basic functions for power analysis.
- Cohen J. (1988). Statistical Power Analysis for the Behavioral Sciences.
Session info
R version 4.4.1 (2024-06-14)
Platform: x86_64-pc-linux-gnu
Running under: Ubuntu 24.04.4 LTS
Matrix products: default
BLAS: /usr/lib/x86_64-linux-gnu/openblas-pthread/libblas.so.3
LAPACK: /usr/lib/x86_64-linux-gnu/openblas-pthread/libopenblasp-r0.3.26.so; LAPACK version 3.12.0
locale:
[1] LC_CTYPE=C.UTF-8 LC_NUMERIC=C LC_TIME=C.UTF-8
[4] LC_COLLATE=C.UTF-8 LC_MONETARY=C.UTF-8 LC_MESSAGES=C.UTF-8
[7] LC_PAPER=C.UTF-8 LC_NAME=C LC_ADDRESS=C
[10] LC_TELEPHONE=C LC_MEASUREMENT=C.UTF-8 LC_IDENTIFICATION=C
time zone: UTC
tzcode source: system (glibc)
attached base packages:
[1] stats graphics grDevices utils datasets methods base
other attached packages:
[1] broom_1.0.12 gtsummary_2.5.0 pwr_1.3-0 lubridate_1.9.5
[5] forcats_1.0.1 stringr_1.6.0 dplyr_1.2.1 purrr_1.2.2
[9] readr_2.2.0 tidyr_1.3.2 tibble_3.3.1 ggplot2_4.0.3
[13] tidyverse_2.0.0
loaded via a namespace (and not attached):
[1] gt_1.3.0 sass_0.4.10 utf8_1.2.6 generics_0.1.4
[5] xml2_1.5.2 stringi_1.8.7 hms_1.1.4 digest_0.6.39
[9] magrittr_2.0.5 evaluate_1.0.5 grid_4.4.1 timechange_0.4.0
[13] RColorBrewer_1.1-3 cards_0.7.1 fastmap_1.2.0 jsonlite_2.0.0
[17] cardx_0.3.2 backports_1.5.1 scales_1.4.0 cli_3.6.6
[21] rlang_1.2.0 litedown_0.9 commonmark_2.0.0 base64enc_0.1-6
[25] withr_3.0.2 yaml_2.3.12 otel_0.2.0 tools_4.4.1
[29] tzdb_0.5.0 vctrs_0.7.3 R6_2.6.1 lifecycle_1.0.5
[33] fs_2.1.0 htmlwidgets_1.6.4 pkgconfig_2.0.3 pillar_1.11.1
[37] gtable_0.3.6 glue_1.8.1 xfun_0.57 tidyselect_1.2.1
[41] knitr_1.51 farver_2.1.2 htmltools_0.5.9 rmarkdown_2.31
[45] labeling_0.4.3 compiler_4.4.1 S7_0.2.2 markdown_2.0