Week 2, Session 1 — One-way ANOVA and contrasts (emmeans)
Course 2 — #courses
Note
Inference labs use the five-step template: Hypothesis → Visualise → Assumptions → Conduct → Conclude.
Learning objectives
- Fit a one-way ANOVA with
aov()and interpret the omnibus F-test. - Use
emmeansto obtain estimated marginal means and pairwise contrasts with controlled family-wise error. - Read a Tukey HSD output and translate it into a sentence.
Prerequisites
Course 1 t-tests; Course 2 Week 1 regression basics.
Background
Analysis of variance is linear regression with a categorical predictor and a tradition. The omnibus F-test asks whether any group means differ; the follow-up contrasts ask which ones. The correct order is omnibus first, then targeted contrasts, preferably with a multiple testing correction that respects the family of tests you intended before seeing the data.
The emmeans package has made the post-hoc machinery substantially cleaner. emmeans(fit, ~ group) returns the estimated marginal means on the response scale; pairs() produces the pairwise differences with Tukey-adjusted p-values by default.
A common mistake is to run pairwise t-tests without correction and report the one with the smallest p-value. A less common but more insidious mistake is to run the omnibus F, see it is significant, and stop — reporting a single p-value without naming which groups differ and by how much.
Setup
1. Hypothesis
Using ToothGrowth, ask whether guinea-pig tooth length depends on dose of vitamin C.
Null: mean tooth length is equal across the three doses. Alternative: at least one dose differs.
2. Visualise
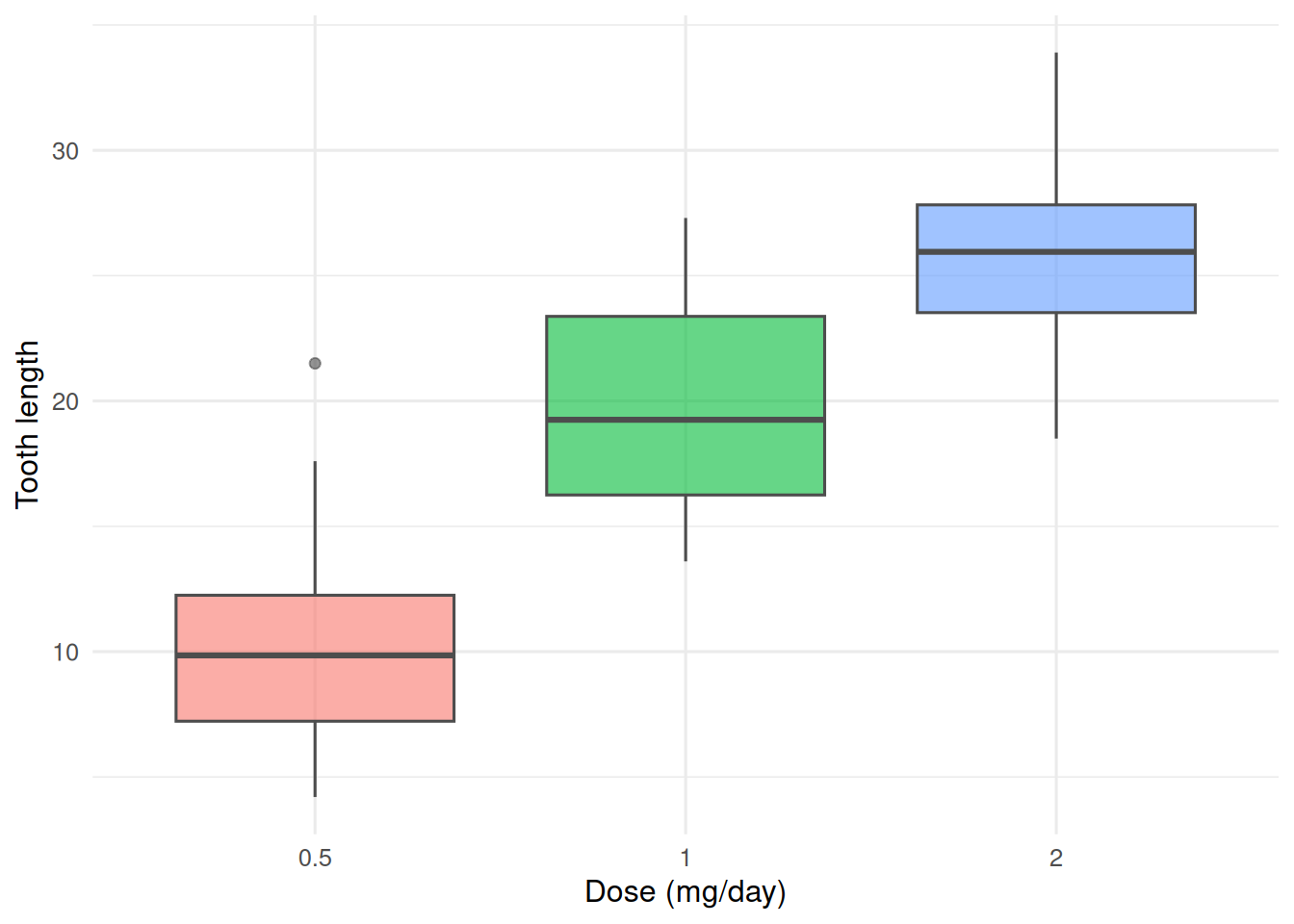
3. Assumptions
Approximate normality within groups and equal variances.
Bartlett test of homogeneity of variances
data: len by dose
Bartlett's K-squared = 0.66547, df = 2, p-value = 0.717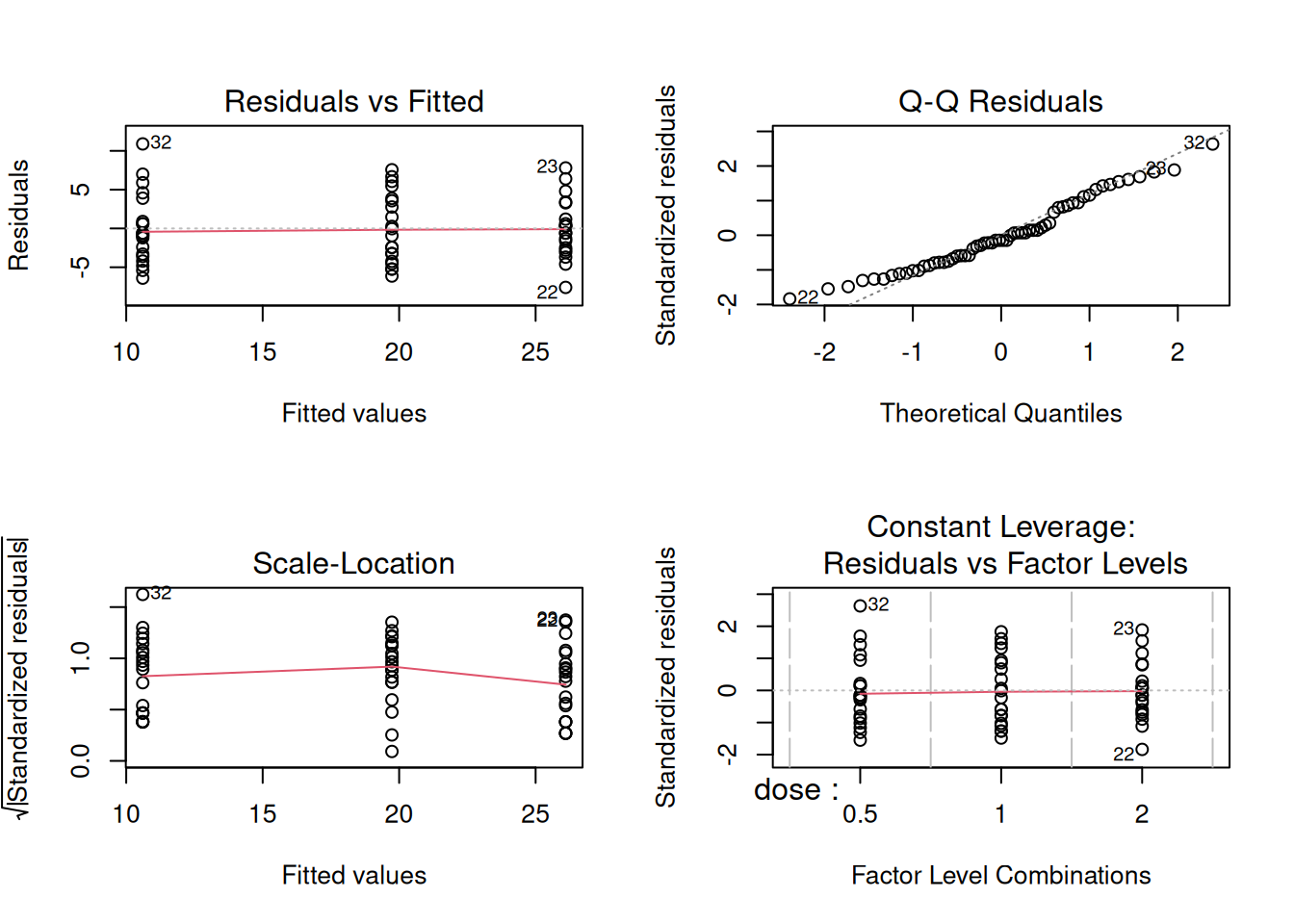
4. Conduct
Df Sum Sq Mean Sq F value Pr(>F)
dose 2 2426 1213 67.42 9.53e-16 ***
Residuals 57 1026 18
---
Signif. codes: 0 '***' 0.001 '**' 0.01 '*' 0.05 '.' 0.1 ' ' 1 dose emmean SE df lower.CL upper.CL
0.5 10.6 0.949 57 8.71 12.5
1 19.7 0.949 57 17.84 21.6
2 26.1 0.949 57 24.20 28.0
Confidence level used: 0.95 contrast estimate SE df t.ratio p.value
dose0.5 - dose1 -9.13 1.34 57 -6.806 <0.0001
dose0.5 - dose2 -15.49 1.34 57 -11.551 <0.0001
dose1 - dose2 -6.37 1.34 57 -4.745 <0.0001
P value adjustment: tukey method for comparing a family of 3 estimates 5. Concluding statement
Tooth length increased with dose of vitamin C (one-way ANOVA F = 67.42; p = 9.5^{-16}). Tukey-adjusted contrasts indicated that the 2 mg/day dose gave longer teeth than both lower doses (all adjusted p < 0.05).
Common pitfalls
- Stopping at a significant F without reporting which contrasts drive it.
- Reporting unadjusted pairwise p-values after the omnibus test.
- Treating a non-significant Bartlett test as proof of equal variances.
Further reading
- Lenth RV. emmeans package vignette.
- Oehlert GW. A First Course in Design and Analysis of Experiments.
- Maxwell SE, Delaney HD. Designing Experiments and Analyzing Data.
Session info
R version 4.4.1 (2024-06-14)
Platform: x86_64-pc-linux-gnu
Running under: Ubuntu 24.04.4 LTS
Matrix products: default
BLAS: /usr/lib/x86_64-linux-gnu/openblas-pthread/libblas.so.3
LAPACK: /usr/lib/x86_64-linux-gnu/openblas-pthread/libopenblasp-r0.3.26.so; LAPACK version 3.12.0
locale:
[1] LC_CTYPE=C.UTF-8 LC_NUMERIC=C LC_TIME=C.UTF-8
[4] LC_COLLATE=C.UTF-8 LC_MONETARY=C.UTF-8 LC_MESSAGES=C.UTF-8
[7] LC_PAPER=C.UTF-8 LC_NAME=C LC_ADDRESS=C
[10] LC_TELEPHONE=C LC_MEASUREMENT=C.UTF-8 LC_IDENTIFICATION=C
time zone: UTC
tzcode source: system (glibc)
attached base packages:
[1] stats graphics grDevices utils datasets methods base
other attached packages:
[1] emmeans_2.0.3 broom_1.0.12 lubridate_1.9.5 forcats_1.0.1
[5] stringr_1.6.0 dplyr_1.2.1 purrr_1.2.2 readr_2.2.0
[9] tidyr_1.3.2 tibble_3.3.1 ggplot2_4.0.3 tidyverse_2.0.0
loaded via a namespace (and not attached):
[1] gtable_0.3.6 jsonlite_2.0.0 compiler_4.4.1 tidyselect_1.2.1
[5] scales_1.4.0 yaml_2.3.12 fastmap_1.2.0 lattice_0.22-6
[9] coda_0.19-4.1 R6_2.6.1 labeling_0.4.3 generics_0.1.4
[13] knitr_1.51 backports_1.5.1 htmlwidgets_1.6.4 pillar_1.11.1
[17] RColorBrewer_1.1-3 tzdb_0.5.0 rlang_1.2.0 stringi_1.8.7
[21] xfun_0.57 S7_0.2.2 otel_0.2.0 estimability_1.5.1
[25] timechange_0.4.0 cli_3.6.6 withr_3.0.2 magrittr_2.0.5
[29] digest_0.6.39 grid_4.4.1 xtable_1.8-8 mvtnorm_1.3-7
[33] hms_1.1.4 lifecycle_1.0.5 vctrs_0.7.3 evaluate_1.0.5
[37] glue_1.8.1 farver_2.1.2 rmarkdown_2.31 tools_4.4.1
[41] pkgconfig_2.0.3 htmltools_0.5.9