Week 2, Session 4 — GAMs with mgcv
Course 2 — #courses
Note
Inference labs use the five-step template: Hypothesis → Visualise → Assumptions → Conduct → Conclude.
Learning objectives
- Fit a generalised additive model with
mgcv::gam()and a single smooth term. - Read effective degrees of freedom as a data-driven measure of flexibility.
- Visualise a smooth with
gratia::draw()and report its shape in plain language.
Prerequisites
Sessions 1–2 of Week 1.
Background
Generalised additive models replace the linear term β·X in a regression with a smooth function f(X) estimated from the data. The smooth is regularised so that wiggliness is penalised; the penalty strength is chosen by restricted maximum likelihood or generalised cross-validation. The effective degrees of freedom (edf) of the smooth quantifies how much of that flexibility was used: an edf near 1 means the smooth is essentially a line; an edf of 5 means something meaningfully curved.
GAMs are particularly useful when the shape of the relationship is the scientific question. They are also useful as diagnostics for linear models, because plotting a smooth against a predictor can reveal a curve that the linear term would miss.
The mgcv package is the reference implementation. gratia is a newer graphics layer on top of mgcv that produces tidy, ggplot-based plots of smooths, partial effects, and derivatives.
Setup
1. Hypothesis
Simulate a sine-wave-plus-noise relationship and ask whether a GAM recovers it while a linear model misses it.
2. Visualise
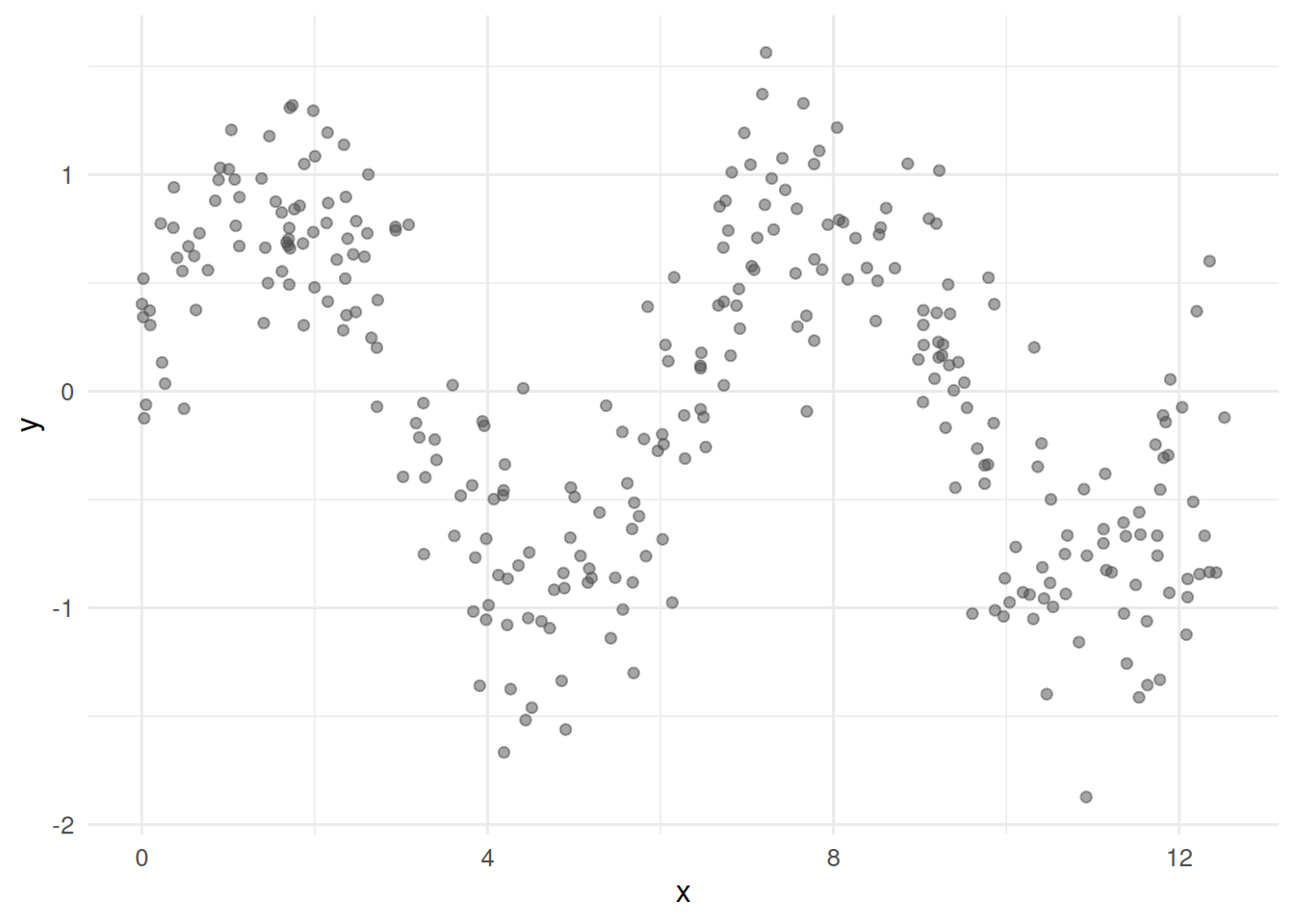
3. Assumptions
GAMs assume independent observations and a correctly specified response distribution. The smooth flexibility is chosen automatically; the user chooses the basis and, if relevant, the upper limit on flexibility via k.
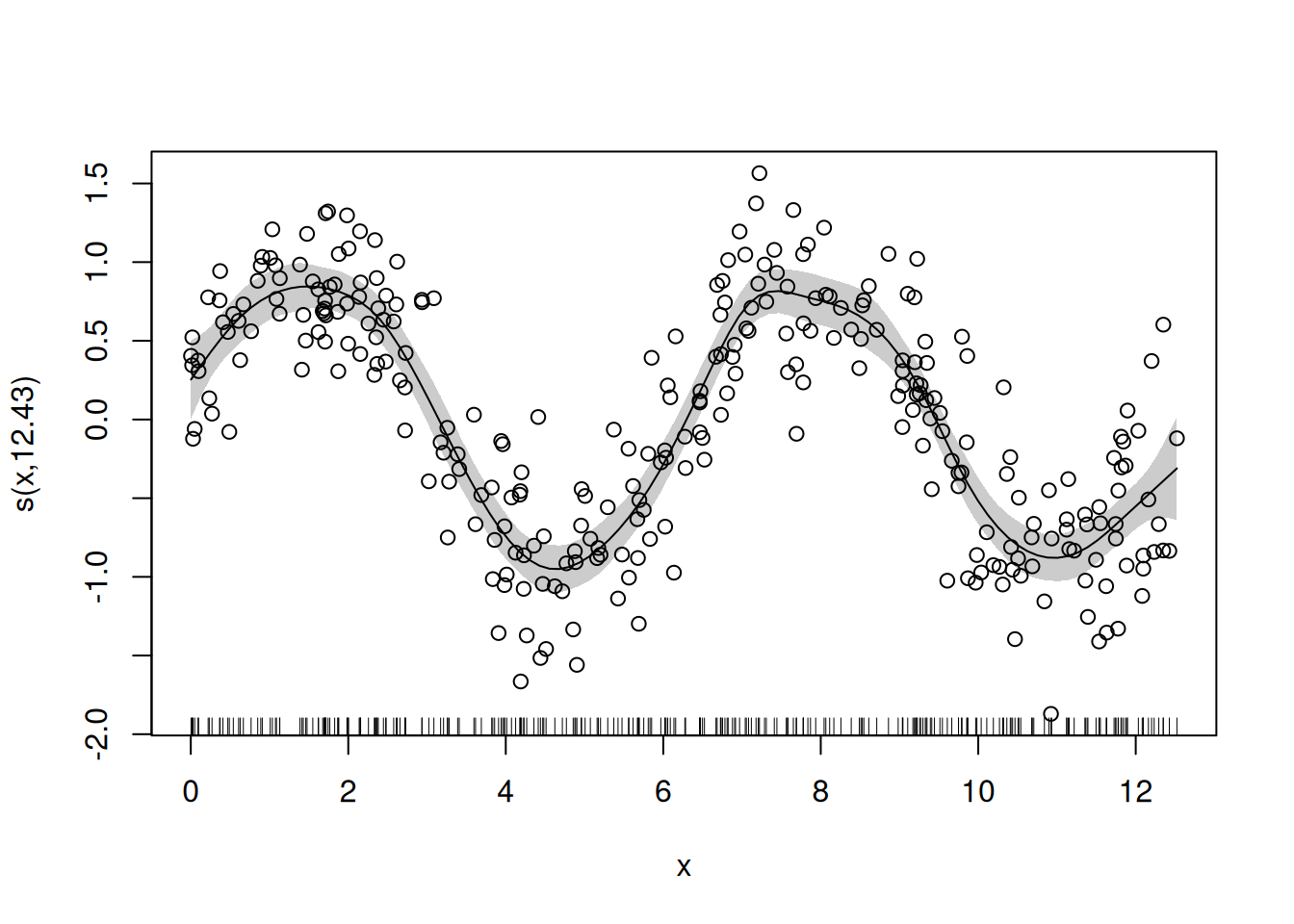
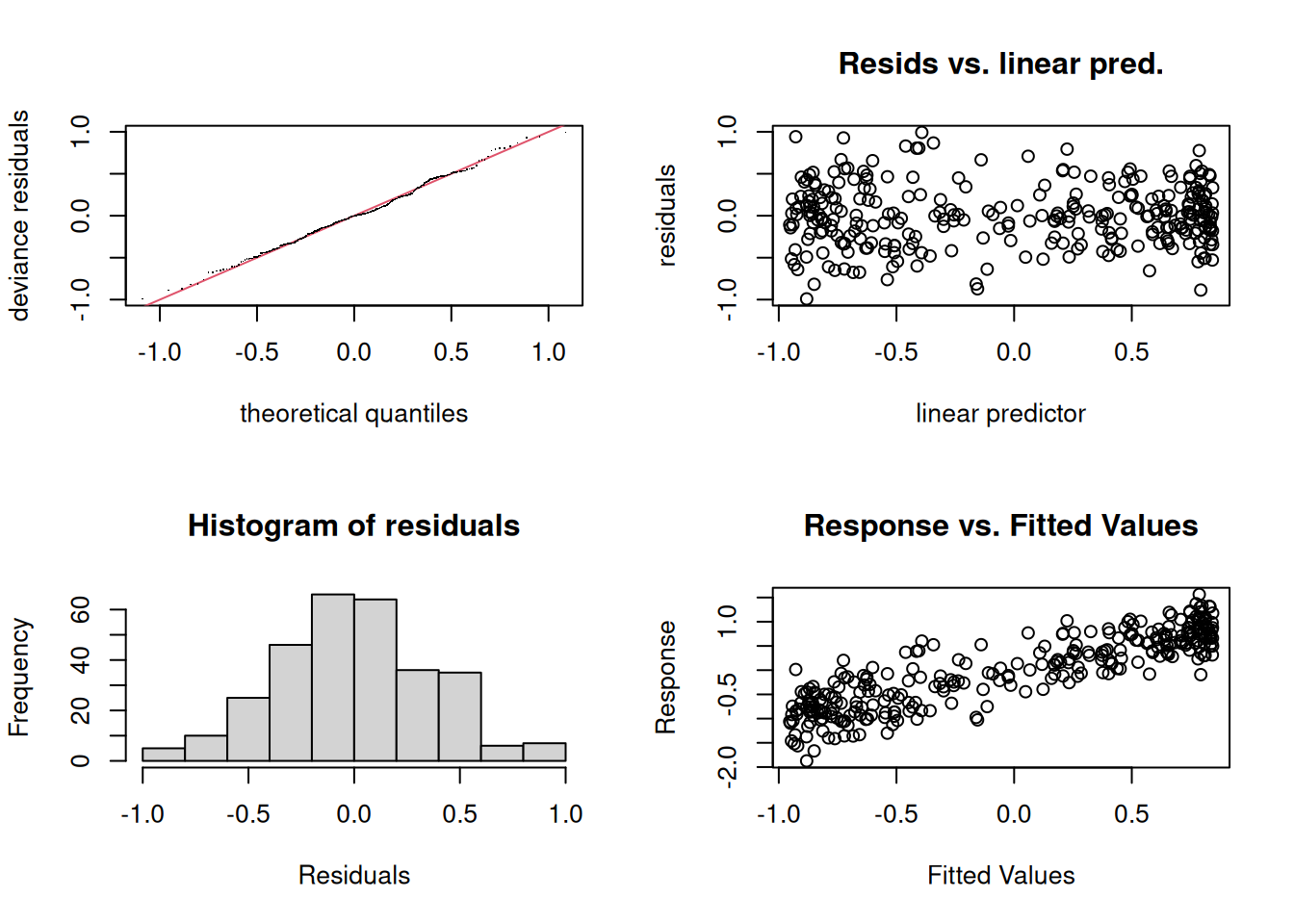
Method: REML Optimizer: outer newton
full convergence after 4 iterations.
Gradient range [-5.838318e-05,1.726219e-06]
(score 150.8396 & scale 0.1374259).
Hessian positive definite, eigenvalue range [4.979807,149.2267].
Model rank = 20 / 20
Basis dimension (k) checking results. Low p-value (k-index<1) may
indicate that k is too low, especially if edf is close to k'.
k' edf k-index p-value
s(x) 19.0 12.4 1.11 0.99gam.check reports k-index and effective df — an indication of whether k is large enough.
4. Conduct
Family: gaussian
Link function: identity
Formula:
y ~ s(x, bs = "cr", k = 20)
Parametric coefficients:
Estimate Std. Error t value Pr(>|t|)
(Intercept) -0.002187 0.021403 -0.102 0.919
Approximate significance of smooth terms:
edf Ref.df F p-value
s(x) 12.43 14.8 62.12 <2e-16 ***
---
Signif. codes: 0 '***' 0.001 '**' 0.01 '*' 0.05 '.' 0.1 ' ' 1
R-sq.(adj) = 0.755 Deviance explained = 76.5%
-REML = 150.84 Scale est. = 0.13743 n = 300 df AIC
fit_lm 3.00000 631.7192
fit_gam 14.69402 271.6137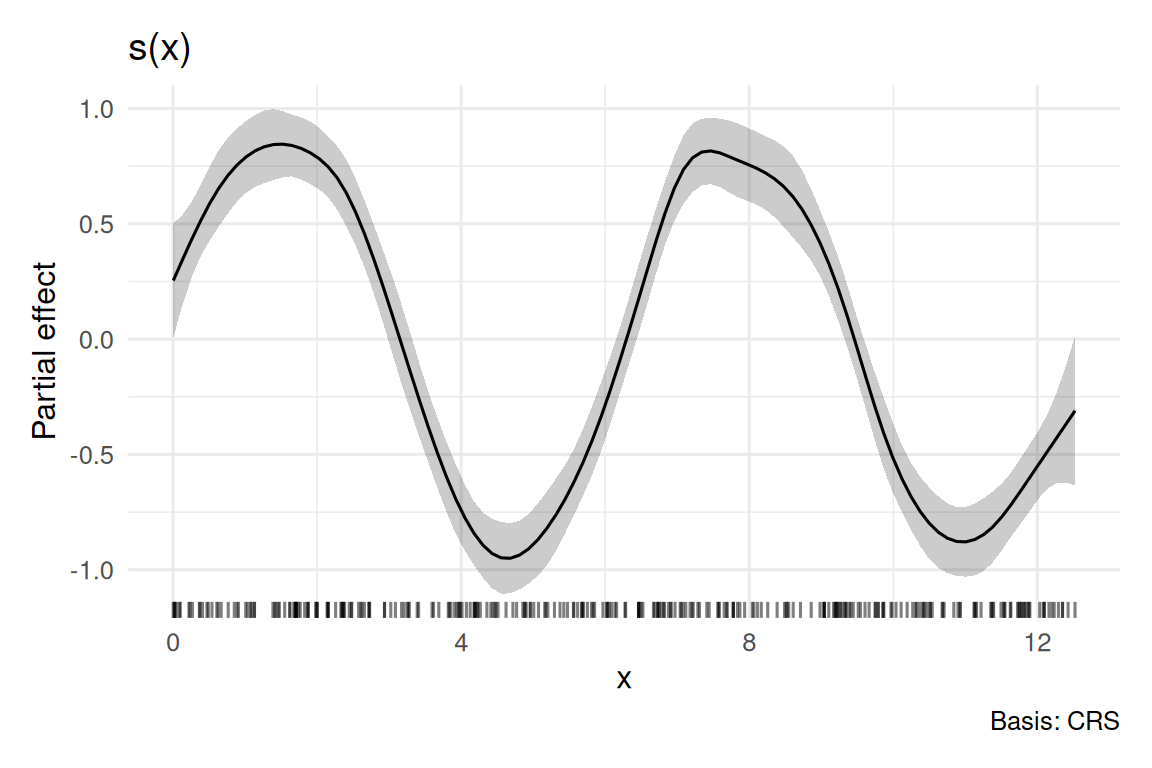
5. Concluding statement
A GAM with a single smooth of x captured the non-linear relationship (edf = 12.4; approximate p < 0.001). A linear model gave a near-zero slope because the underlying function is a full sine wave. The GAM is preferred when the shape, not a scalar slope, is the scientific object.
Common pitfalls
- Setting
ktoo small and constraining the smooth. - Comparing GAMs with AIC as if it were a likelihood-ratio test.
- Reporting only the smooth without showing its plot.
Further reading
- Wood SN. Generalized Additive Models: An Introduction with R.
- Pedersen EJ et al. (2019), Hierarchical generalized additive models.
- Simpson GL. gratia package vignette.
Session info
R version 4.4.1 (2024-06-14)
Platform: x86_64-pc-linux-gnu
Running under: Ubuntu 24.04.4 LTS
Matrix products: default
BLAS: /usr/lib/x86_64-linux-gnu/openblas-pthread/libblas.so.3
LAPACK: /usr/lib/x86_64-linux-gnu/openblas-pthread/libopenblasp-r0.3.26.so; LAPACK version 3.12.0
locale:
[1] LC_CTYPE=C.UTF-8 LC_NUMERIC=C LC_TIME=C.UTF-8
[4] LC_COLLATE=C.UTF-8 LC_MONETARY=C.UTF-8 LC_MESSAGES=C.UTF-8
[7] LC_PAPER=C.UTF-8 LC_NAME=C LC_ADDRESS=C
[10] LC_TELEPHONE=C LC_MEASUREMENT=C.UTF-8 LC_IDENTIFICATION=C
time zone: UTC
tzcode source: system (glibc)
attached base packages:
[1] stats graphics grDevices utils datasets methods base
other attached packages:
[1] broom_1.0.12 gratia_0.11.2 mgcv_1.9-1 nlme_3.1-164
[5] lubridate_1.9.5 forcats_1.0.1 stringr_1.6.0 dplyr_1.2.1
[9] purrr_1.2.2 readr_2.2.0 tidyr_1.3.2 tibble_3.3.1
[13] ggplot2_4.0.3 tidyverse_2.0.0
loaded via a namespace (and not attached):
[1] generics_0.1.4 stringi_1.8.7 lattice_0.22-6 hms_1.1.4
[5] digest_0.6.39 magrittr_2.0.5 evaluate_1.0.5 grid_4.4.1
[9] timechange_0.4.0 RColorBrewer_1.1-3 fastmap_1.2.0 jsonlite_2.0.0
[13] Matrix_1.7-0 backports_1.5.1 ggokabeito_0.1.0 scales_1.4.0
[17] cli_3.6.6 rlang_1.2.0 splines_4.4.1 withr_3.0.2
[21] yaml_2.3.12 otel_0.2.0 tools_4.4.1 tzdb_0.5.0
[25] nanonext_1.9.0 vctrs_0.7.3 R6_2.6.1 lifecycle_1.0.5
[29] tweedie_3.0.19 htmlwidgets_1.6.4 pkgconfig_2.0.3 pillar_1.11.1
[33] gtable_0.3.6 glue_1.8.1 Rcpp_1.1.1-1.1 statmod_1.5.1
[37] xfun_0.57 tidyselect_1.2.1 knitr_1.51 farver_2.1.2
[41] patchwork_1.3.2 htmltools_0.5.9 mirai_2.6.1 labeling_0.4.3
[45] rmarkdown_2.31 compiler_4.4.1 S7_0.2.2 mvnfast_0.2.8