Week 3, Session 3 — Ordinal and multinomial regression
Course 2 — #courses
Note
Inference labs use the five-step template: Hypothesis → Visualise → Assumptions → Conduct → Conclude.
Learning objectives
- Fit a proportional-odds model with
MASS::polr()and report a cumulative odds ratio. - Fit a multinomial logistic regression with
nnet::multinom()for an unordered outcome. - Check the proportional-odds assumption and decide what to do when it fails.
Prerequisites
Week 3 Session 1 on binary logistic regression.
Background
Ordinal outcomes — pain scales, tumour stages, Likert items — carry more information than a dichotomised version and usually less than a continuous version. The proportional-odds model (polr) assumes the effect of each predictor is the same across all cumulative splits of the response; the single coefficient is the log cumulative odds ratio. Multinomial logistic regression makes no such constraint and fits a separate logit for each non-reference category.
The proportional-odds assumption is strong. A Brant test (in the brant package) or simply comparing the polr fit to a multinomial fit on deviance can detect violations. When it fails, options include a partial proportional-odds model, the multinomial model, or a continuation-ratio model.
Multinomial models are notoriously hungry for data: each non-reference category requires a full set of coefficients. Reporting should list all coefficient tables and a global test (likelihood ratio) rather than a single p-value.
Setup
1. Hypothesis
Simulate an ordinal outcome (symptom severity in three levels) driven by a single continuous exposure, then contrast the ordinal and multinomial analyses.
2. Visualise
n <- 400
x <- rnorm(n)
eta <- 1.2 * x
# cumulative cutpoints
probs <- cbind(
plogis(-0.5 - eta),
plogis(1.0 - eta) - plogis(-0.5 - eta),
1 - plogis(1.0 - eta)
)
y <- apply(probs, 1, function(p) sample(1:3, 1, prob = p))
dat <- tibble(x, y = factor(y, levels = 1:3,
labels = c("mild", "moderate", "severe"),
ordered = TRUE))
ggplot(dat, aes(y, x, fill = y)) +
geom_boxplot(alpha = 0.6, colour = "grey30") +
labs(x = "Severity", y = "Exposure x") +
theme(legend.position = "none")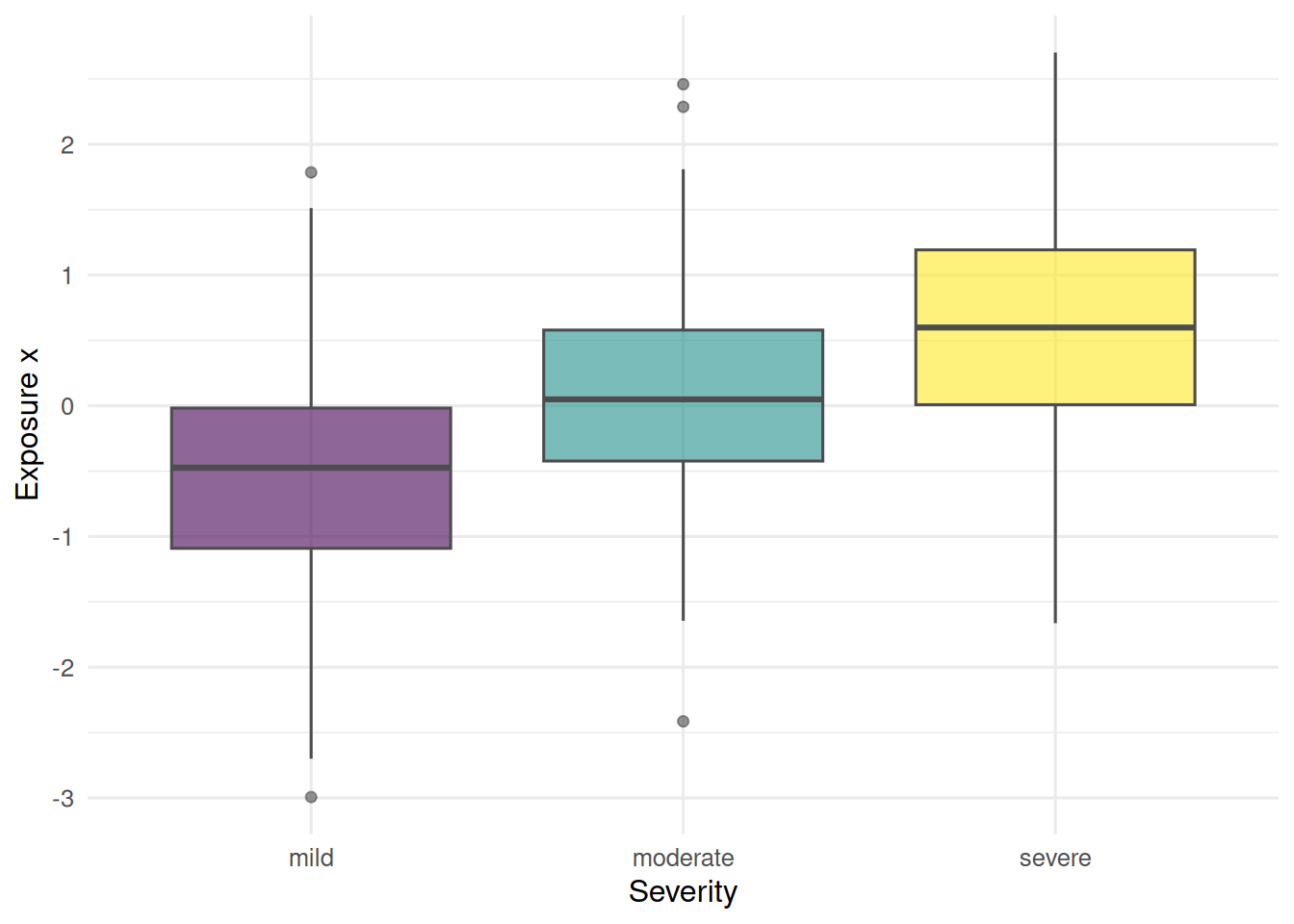
3. Assumptions
Proportional-odds (same β across cumulative splits) for polr; independent observations for both models.
# quick graphical check: fit logistic at each cumulative split and compare slopes
splits <- tibble(cut = c("mild vs rest", "mild+mod vs severe")) |>
mutate(coef = c(
coef(glm(I(as.numeric(y) >= 2) ~ x, data = dat, family = binomial))[2],
coef(glm(I(as.numeric(y) >= 3) ~ x, data = dat, family = binomial))[2]
))
splits# A tibble: 2 × 2
cut coef
<chr> <dbl>
1 mild vs rest 1.18
2 mild+mod vs severe 1.16If the two slopes are close, proportional odds is plausible.
4. Conduct
Call:
polr(formula = y ~ x, data = dat, Hess = TRUE)
Coefficients:
Value Std. Error t value
x 1.162 0.1249 9.297
Intercepts:
Value Std. Error t value
mild|moderate -0.4652 0.1127 -4.1261
moderate|severe 0.9053 0.1198 7.5569
Residual Deviance: 757.0974
AIC: 763.0974 Value Std. Error t value p value
x 1.1616521 0.1249448 9.297319 0
mild|moderate -0.4651740 0.1127396 -4.126092 0
moderate|severe 0.9053007 0.1197974 7.556932 0 x
3.195208 Multinomial:
Call:
multinom(formula = y ~ x, data = dat, trace = FALSE)
Coefficients:
(Intercept) x
moderate -0.2454994 0.8399669
severe -0.2768431 1.5766088
Std. Errors:
(Intercept) x
moderate 0.1348679 0.1638624
severe 0.1417515 0.1815418
Residual Deviance: 756.7975
AIC: 764.7975 (Intercept) x
moderate 0.069 0
severe 0.051 05. Concluding statement
The proportional-odds model returned a cumulative odds ratio per unit of x of 3.2, meaning larger x shifted symptom severity upward. The multinomial model returned two logits (moderate vs mild and severe vs mild) with effects in the same direction; deviance-based comparison did not reject the proportional-odds simplification.
Common pitfalls
- Reporting only the odds ratio and ignoring whether proportional-odds holds.
- Using a multinomial model with a tiny reference category.
- Treating an ordered outcome as continuous and fitting a linear model.
Further reading
- Agresti A. Analysis of Ordinal Categorical Data.
- Harrell FE. Regression Modeling Strategies, ch. 13.
- Venables WN, Ripley BD. Modern Applied Statistics with S, ch. 7.
Session info
R version 4.4.1 (2024-06-14)
Platform: x86_64-pc-linux-gnu
Running under: Ubuntu 24.04.4 LTS
Matrix products: default
BLAS: /usr/lib/x86_64-linux-gnu/openblas-pthread/libblas.so.3
LAPACK: /usr/lib/x86_64-linux-gnu/openblas-pthread/libopenblasp-r0.3.26.so; LAPACK version 3.12.0
locale:
[1] LC_CTYPE=C.UTF-8 LC_NUMERIC=C LC_TIME=C.UTF-8
[4] LC_COLLATE=C.UTF-8 LC_MONETARY=C.UTF-8 LC_MESSAGES=C.UTF-8
[7] LC_PAPER=C.UTF-8 LC_NAME=C LC_ADDRESS=C
[10] LC_TELEPHONE=C LC_MEASUREMENT=C.UTF-8 LC_IDENTIFICATION=C
time zone: UTC
tzcode source: system (glibc)
attached base packages:
[1] stats graphics grDevices utils datasets methods base
other attached packages:
[1] nnet_7.3-19 MASS_7.3-60.2 broom_1.0.12 lubridate_1.9.5
[5] forcats_1.0.1 stringr_1.6.0 dplyr_1.2.1 purrr_1.2.2
[9] readr_2.2.0 tidyr_1.3.2 tibble_3.3.1 ggplot2_4.0.3
[13] tidyverse_2.0.0
loaded via a namespace (and not attached):
[1] gtable_0.3.6 jsonlite_2.0.0 compiler_4.4.1 tidyselect_1.2.1
[5] scales_1.4.0 yaml_2.3.12 fastmap_1.2.0 R6_2.6.1
[9] labeling_0.4.3 generics_0.1.4 knitr_1.51 backports_1.5.1
[13] htmlwidgets_1.6.4 pillar_1.11.1 RColorBrewer_1.1-3 tzdb_0.5.0
[17] rlang_1.2.0 utf8_1.2.6 stringi_1.8.7 xfun_0.57
[21] S7_0.2.2 otel_0.2.0 viridisLite_0.4.3 timechange_0.4.0
[25] cli_3.6.6 withr_3.0.2 magrittr_2.0.5 digest_0.6.39
[29] grid_4.4.1 hms_1.1.4 lifecycle_1.0.5 vctrs_0.7.3
[33] evaluate_1.0.5 glue_1.8.1 farver_2.1.2 rmarkdown_2.31
[37] tools_4.4.1 pkgconfig_2.0.3 htmltools_0.5.9