Week 1, Session 2 — Randomised controlled trials
Course 3 — #courses
Note
Inference lab using the five-step template: Hypothesis → Visualise → Assumptions → Conduct → Conclude.
Learning objectives
- Contrast parallel-group, crossover, cluster, and factorial RCT designs.
- Simulate a parallel-group RCT with a realistic allocation procedure.
- Interpret intention-to-treat (ITT) and per-protocol (PP) analyses and say why they can differ.
Prerequisites
Course 1 inference; t.test(), lm(), and basic tidyverse.
Background
The randomised controlled trial (RCT) is the reference design for causal inference because randomisation balances both measured and unmeasured confounders in expectation. Parallel-group RCTs assign each participant to one arm for the duration of the study. Crossover trials give each participant every treatment in a randomised order and rely on a washout period. Cluster trials randomise groups (clinics, villages) rather than individuals. Factorial trials vary more than one treatment at once and can answer more than one question per patient.
Every RCT has two analyses that will often disagree. Intention-to- treat analyses everyone in the arm they were randomised to, regardless of what they actually received. Per-protocol analyses only those who received the assigned treatment as planned. ITT is conservative for superiority and preserves the randomisation; PP is informative for efficacy in compliant patients but can be biased. Report both, and say which is primary in the protocol.
Allocation concealment — the process that keeps the next assignment unknown until the patient has been enrolled — is not the same as blinding. You can have one without the other, and you need both for the trial to protect itself from selection bias and differential outcome ascertainment.
Setup
1. Hypothesis
Treatment reduces the outcome (a continuous symptom score) relative to placebo. We will also see what happens when 15% of participants cross over from treatment to placebo after randomisation.
2. Visualise
n <- 200
trial <- tibble(
id = seq_len(n),
arm = sample(rep(c("placebo", "treatment"), each = n / 2)),
# 15% of treatment arm never take the drug (will be placebo in reality)
crossed = if_else(arm == "treatment" & runif(n) < 0.15, TRUE, FALSE),
received = if_else(crossed, "placebo", arm),
y = rnorm(n, mean = 50, sd = 8) +
if_else(received == "treatment", -5, 0)
)
trial |>
ggplot(aes(arm, y, fill = arm)) +
geom_boxplot(alpha = 0.6, colour = "grey30") +
labs(x = NULL, y = "Symptom score") +
theme(legend.position = "none")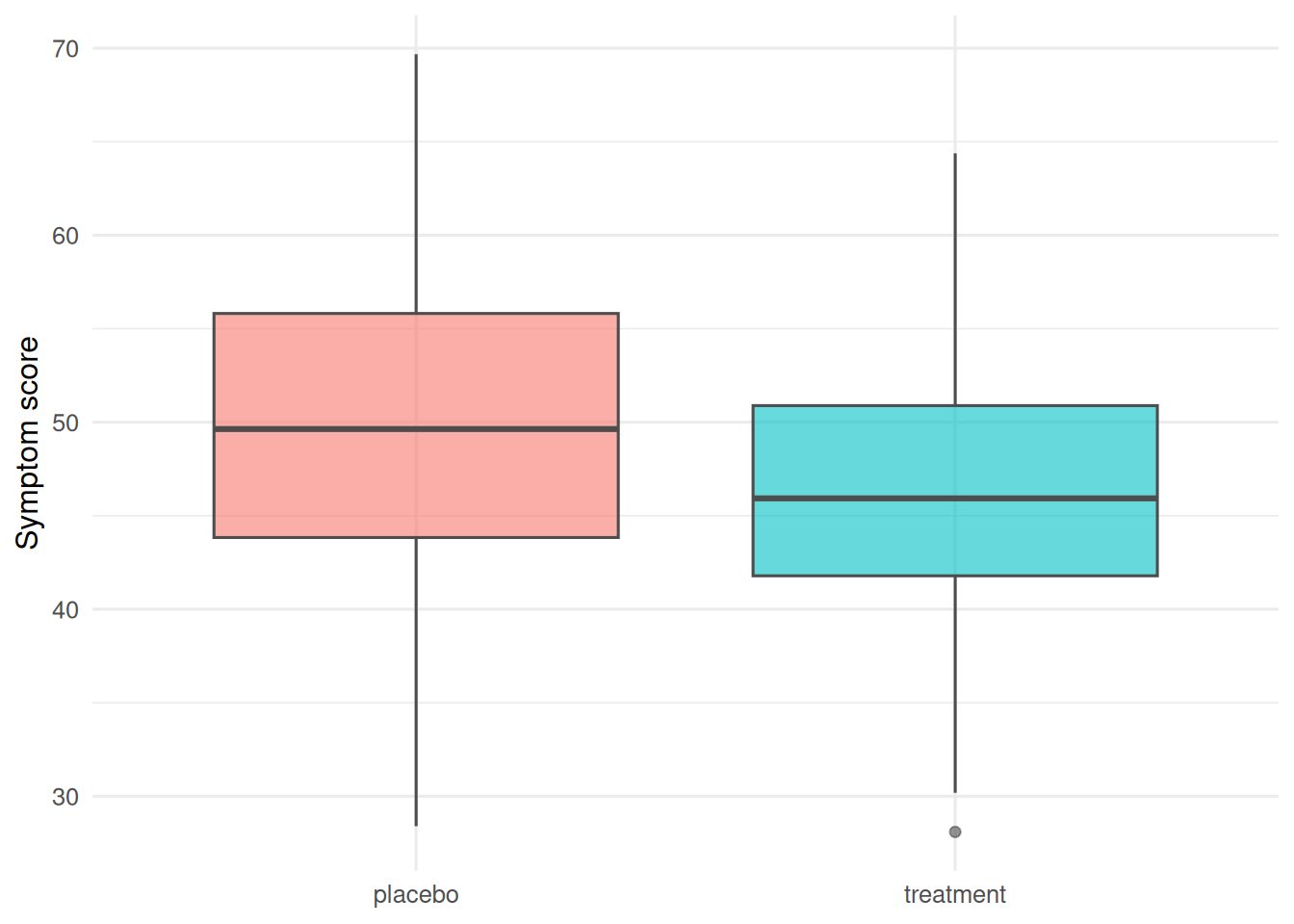
3. Assumptions
A two-sample t-test on the ITT population assumes independent observations within arms and roughly normal residuals within each arm. We do not assume equal variances; t.test() uses Welch by default. The more important assumption is that randomisation worked — that the allocation is independent of every baseline covariate.
4. Conduct
Welch Two Sample t-test
data: y by arm
t = 3.3683, df = 194.51, p-value = 0.0009118
alternative hypothesis: true difference in means between group placebo and group treatment is not equal to 0
95 percent confidence interval:
1.508930 5.772362
sample estimates:
mean in group placebo mean in group treatment
49.69820 46.05756
Welch Two Sample t-test
data: y by received
t = 3.4212, df = 173.54, p-value = 0.0007774
alternative hypothesis: true difference in means between group placebo and group treatment is not equal to 0
95 percent confidence interval:
1.580709 5.891547
sample estimates:
mean in group placebo mean in group treatment
49.35365 45.61752 The ITT estimate is closer to zero than the PP estimate because the crossed-over patients pull the treatment arm mean toward placebo.
5. Concluding statement
In a simulated parallel-group RCT (n = 200), the ITT analysis showed a mean difference of 3.6 (95% CI: -5.8 to -1.5) on the symptom score; a per-protocol analysis gave a larger estimated benefit of 3.7, illustrating the usual direction of disagreement when non-adherence is informative.
Common pitfalls
- Calling a non-random allocation “randomised” because a coin was flipped after enrolment.
- Reporting only PP and calling it primary.
- Ignoring stratification variables in the analysis when they were used at randomisation.
- Cluster trials analysed as if individuals were independent.
Further reading
- Schulz KF, Altman DG, Moher D (2010), CONSORT 2010 Statement.
- Pocock SJ. Clinical Trials: A Practical Approach.
- Hernán MA, Hernández-Díaz S (2012), Beyond the intention-to-treat in comparative effectiveness research.
Session info
R version 4.4.1 (2024-06-14)
Platform: x86_64-pc-linux-gnu
Running under: Ubuntu 24.04.4 LTS
Matrix products: default
BLAS: /usr/lib/x86_64-linux-gnu/openblas-pthread/libblas.so.3
LAPACK: /usr/lib/x86_64-linux-gnu/openblas-pthread/libopenblasp-r0.3.26.so; LAPACK version 3.12.0
locale:
[1] LC_CTYPE=C.UTF-8 LC_NUMERIC=C LC_TIME=C.UTF-8
[4] LC_COLLATE=C.UTF-8 LC_MONETARY=C.UTF-8 LC_MESSAGES=C.UTF-8
[7] LC_PAPER=C.UTF-8 LC_NAME=C LC_ADDRESS=C
[10] LC_TELEPHONE=C LC_MEASUREMENT=C.UTF-8 LC_IDENTIFICATION=C
time zone: UTC
tzcode source: system (glibc)
attached base packages:
[1] stats graphics grDevices utils datasets methods base
other attached packages:
[1] lubridate_1.9.5 forcats_1.0.1 stringr_1.6.0 dplyr_1.2.1
[5] purrr_1.2.2 readr_2.2.0 tidyr_1.3.2 tibble_3.3.1
[9] ggplot2_4.0.3 tidyverse_2.0.0
loaded via a namespace (and not attached):
[1] gtable_0.3.6 jsonlite_2.0.0 compiler_4.4.1 tidyselect_1.2.1
[5] scales_1.4.0 yaml_2.3.12 fastmap_1.2.0 R6_2.6.1
[9] labeling_0.4.3 generics_0.1.4 knitr_1.51 htmlwidgets_1.6.4
[13] pillar_1.11.1 RColorBrewer_1.1-3 tzdb_0.5.0 rlang_1.2.0
[17] utf8_1.2.6 stringi_1.8.7 xfun_0.57 S7_0.2.2
[21] otel_0.2.0 timechange_0.4.0 cli_3.6.6 withr_3.0.2
[25] magrittr_2.0.5 digest_0.6.39 grid_4.4.1 hms_1.1.4
[29] lifecycle_1.0.5 vctrs_0.7.3 evaluate_1.0.5 glue_1.8.1
[33] farver_2.1.2 rmarkdown_2.31 tools_4.4.1 pkgconfig_2.0.3
[37] htmltools_0.5.9