Week 2, Session 3 — Linear mixed models with lme4
Course 3 — #courses
Note
Inference lab using the five-step template: Hypothesis → Visualise → Assumptions → Conduct → Conclude.
Learning objectives
- Fit random-intercept and random-slope linear mixed models to longitudinal data.
- Test whether adding a random slope improves fit (likelihood ratio test with ML).
- Interpret REML estimates of variance components.
Prerequisites
Course 2 regression; familiarity with lm().
Background
Longitudinal data have two sources of variation: differences between people and differences over time within a person. A random-intercept model lets each person have their own average level; a random-slope model lets each person also have their own rate of change. When the slopes genuinely vary, omitting the random slope underestimates the standard error of the fixed-effect slope and produces p-values that are too small.
REML (restricted maximum likelihood) is the default because it gives unbiased estimates of the variance components by integrating out the fixed effects. It is the right choice for reporting variance components. For comparing models that differ in fixed effects, you must refit with ML (likelihood ratio tests on REML fits are invalid); for comparing models that differ only in random effects, REML is fine.
A common misstep is to interpret the variance of the random intercept as the “within-subject variance”. It is the between- subject variance of the intercept. The within-subject variance is the residual variance, σ², once the random effects are accounted for.
Setup
1. Hypothesis
In the sleepstudy data, reaction time increases with days of sleep deprivation, and the rate of increase varies by subject.
2. Visualise
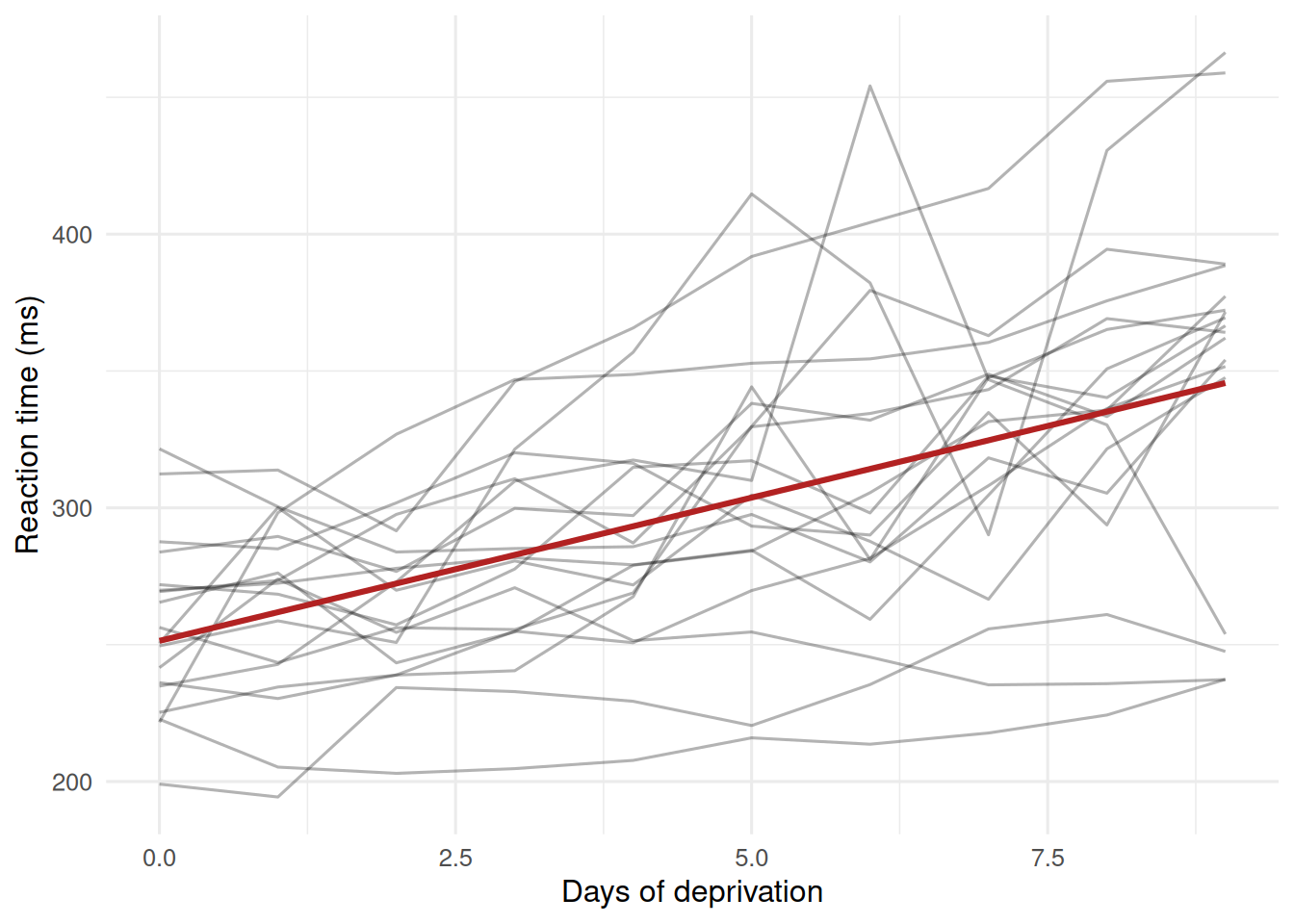
3. Assumptions
Linear relationship between Days and Reaction for each subject; normally distributed random intercepts and slopes (if included); residuals independent within and across subjects after random effects account for clustering; approximately constant residual variance.
4. Conduct
Linear mixed model fit by REML. t-tests use Satterthwaite's method [
lmerModLmerTest]
Formula: Reaction ~ Days + (Days | Subject)
Data: sleepstudy
REML criterion at convergence: 1743.6
Scaled residuals:
Min 1Q Median 3Q Max
-3.9536 -0.4634 0.0231 0.4634 5.1793
Random effects:
Groups Name Variance Std.Dev. Corr
Subject (Intercept) 612.10 24.741
Days 35.07 5.922 0.07
Residual 654.94 25.592
Number of obs: 180, groups: Subject, 18
Fixed effects:
Estimate Std. Error df t value Pr(>|t|)
(Intercept) 251.405 6.825 17.000 36.838 < 2e-16 ***
Days 10.467 1.546 17.000 6.771 3.26e-06 ***
---
Signif. codes: 0 '***' 0.001 '**' 0.01 '*' 0.05 '.' 0.1 ' ' 1
Correlation of Fixed Effects:
(Intr)
Days -0.138Data: sleepstudy
Models:
ri: Reaction ~ Days + (1 | Subject)
rs: Reaction ~ Days + (Days | Subject)
npar AIC BIC logLik -2*log(L) Chisq Df Pr(>Chisq)
ri 4 1794.5 1807.2 -893.23 1786.5
rs 6 1755.6 1774.8 -871.81 1743.6 42.837 2 4.99e-10 ***
---
Signif. codes: 0 '***' 0.001 '**' 0.01 '*' 0.05 '.' 0.1 ' ' 1Linear mixed model fit by maximum likelihood ['lmerModLmerTest']
Formula: Reaction ~ Days + (Days | Subject)
Data: sleepstudy
AIC BIC logLik -2*log(L) df.resid
1763.9393 1783.0971 -875.9697 1751.9393 174
Random effects:
Groups Name Std.Dev. Corr
Subject (Intercept) 23.780
Days 5.717 0.08
Residual 25.592
Number of obs: 180, groups: Subject, 18
Fixed Effects:
(Intercept) Days
251.41 10.47 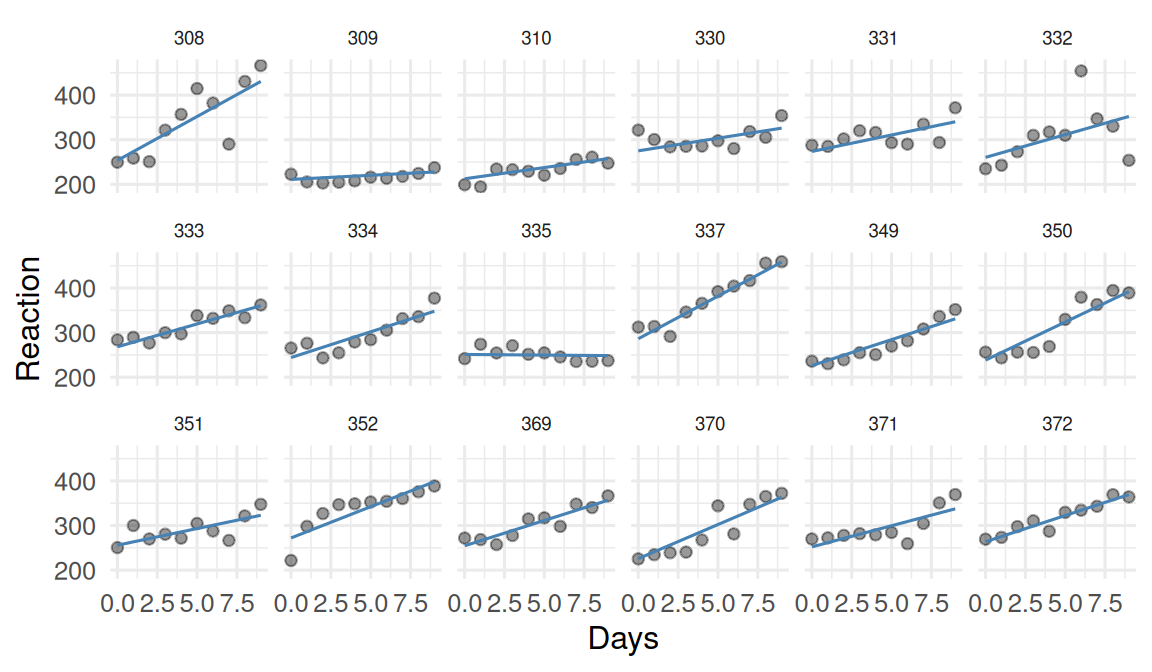
5. Concluding statement
In the sleepstudy data, reaction time increased by about 10.5 ms per day of deprivation, with substantial subject-to-subject variation in slope (SD ≈ 5.9 ms/day). Adding a random slope significantly improved fit over a random-intercept model (likelihood-ratio test p < 0.001).
Common pitfalls
- Testing random-effects with REML-based LRTs without refitting.
- Ignoring the random slope when slopes clearly vary.
- Treating the
Interceptvariance as the within-subject error. - Using p-values from the Wald z on small samples.
Further reading
- Bates D et al. (2015), Fitting linear mixed-effects models using lme4.
- Pinheiro JC, Bates DM. Mixed-Effects Models in S and S-PLUS.
- Gelman A, Hill J. Data Analysis Using Regression and Multilevel/Hierarchical Models.
Session info
R version 4.4.1 (2024-06-14)
Platform: x86_64-pc-linux-gnu
Running under: Ubuntu 24.04.4 LTS
Matrix products: default
BLAS: /usr/lib/x86_64-linux-gnu/openblas-pthread/libblas.so.3
LAPACK: /usr/lib/x86_64-linux-gnu/openblas-pthread/libopenblasp-r0.3.26.so; LAPACK version 3.12.0
locale:
[1] LC_CTYPE=C.UTF-8 LC_NUMERIC=C LC_TIME=C.UTF-8
[4] LC_COLLATE=C.UTF-8 LC_MONETARY=C.UTF-8 LC_MESSAGES=C.UTF-8
[7] LC_PAPER=C.UTF-8 LC_NAME=C LC_ADDRESS=C
[10] LC_TELEPHONE=C LC_MEASUREMENT=C.UTF-8 LC_IDENTIFICATION=C
time zone: UTC
tzcode source: system (glibc)
attached base packages:
[1] stats graphics grDevices utils datasets methods base
other attached packages:
[1] lmerTest_3.2-1 lme4_2.0-1 Matrix_1.7-0 lubridate_1.9.5
[5] forcats_1.0.1 stringr_1.6.0 dplyr_1.2.1 purrr_1.2.2
[9] readr_2.2.0 tidyr_1.3.2 tibble_3.3.1 ggplot2_4.0.3
[13] tidyverse_2.0.0
loaded via a namespace (and not attached):
[1] generics_0.1.4 stringi_1.8.7 lattice_0.22-6
[4] hms_1.1.4 digest_0.6.39 magrittr_2.0.5
[7] evaluate_1.0.5 grid_4.4.1 timechange_0.4.0
[10] RColorBrewer_1.1-3 fastmap_1.2.0 jsonlite_2.0.0
[13] mgcv_1.9-1 scales_1.4.0 numDeriv_2016.8-1.1
[16] reformulas_0.4.4 Rdpack_2.6.6 cli_3.6.6
[19] rlang_1.2.0 rbibutils_2.4.1 splines_4.4.1
[22] withr_3.0.2 yaml_2.3.12 otel_0.2.0
[25] tools_4.4.1 tzdb_0.5.0 nloptr_2.2.1
[28] minqa_1.2.8 boot_1.3-30 vctrs_0.7.3
[31] R6_2.6.1 lifecycle_1.0.5 htmlwidgets_1.6.4
[34] MASS_7.3-60.2 pkgconfig_2.0.3 pillar_1.11.1
[37] gtable_0.3.6 glue_1.8.1 Rcpp_1.1.1-1.1
[40] xfun_0.57 tidyselect_1.2.1 knitr_1.51
[43] farver_2.1.2 htmltools_0.5.9 nlme_3.1-164
[46] labeling_0.4.3 rmarkdown_2.31 compiler_4.4.1
[49] S7_0.2.2