Week 3, Session 2 — Competing risks and multistate models
Course 3 — #courses
Note
Inference lab using the five-step template: Hypothesis → Visualise → Assumptions → Conduct → Conclude.
Learning objectives
- Estimate cause-specific cumulative incidence with
tidycmprsk. - Contrast cause-specific hazards with the Fine-Gray subdistribution hazard.
- Set up a three-state illness-death multistate model with
mstate.
Prerequisites
Course 2 survival analysis.
Background
Competing risks are events that preclude the event of interest: death from cancer competes with death from other causes. Kaplan- Meier treats the competing event as censoring, which is wrong when the censoring is informative. Cumulative incidence functions (CIFs) give the probability that the event of interest has occurred by time t in the presence of competing risks.
Two hazard models are common. Cause-specific Cox models treat each cause in turn, censoring at other causes; they answer an aetiologic question (“does the covariate affect the rate of this cause?”). The Fine-Gray subdistribution hazard model fixes subjects with competing events in the risk set indefinitely; it answers a prognostic question (“does the covariate affect the cumulative incidence?”).
Multistate models generalise survival to several possible transitions. The illness-death model has three states: healthy, ill, dead, with transitions healthy → ill, healthy → dead, and ill → dead. mstate builds these objects from tabular data.
Fine-Gray is descriptive; cause-specific Cox is causal (in the simple, aetiological sense). A covariate can increase the Fine-Gray subdistribution hazard of cause 1 either by increasing cause 1 itself or by decreasing cause 2 — it is a net effect on the cumulative incidence.
Setup
1. Hypothesis
In the survival::colon data (recurrence and death as competing events), a treatment covariate affects both events. We estimate the cumulative incidence of each.
2. Visualise
data(colon, package = "survival")
# Keep death records (etype == 2) + recurrence merged; for teaching
# we build a two-event dataset:
dat <- colon |>
filter(etype == 1) |> # recurrence row per subject
transmute(id, time, status_rec = status,
rx, age, sex) |>
left_join(
colon |> filter(etype == 2) |>
transmute(id, time_d = time, status_d = status),
by = "id"
) |>
mutate(
# event type: 0 censored, 1 recurrence, 2 death without recurrence
etype = case_when(
status_rec == 1 ~ 1L,
status_d == 1 ~ 2L,
TRUE ~ 0L
),
etime = pmin(time, time_d, na.rm = TRUE),
etype_f = factor(etype, levels = 0:2,
labels = c("censored", "recurrence", "death"))
) |>
drop_na(etime, etype_f, rx)
ggplot(dat, aes(etime, fill = etype_f)) +
geom_histogram(bins = 40, alpha = 0.7, position = "identity") +
labs(x = "Time (days)", y = "Count", fill = NULL)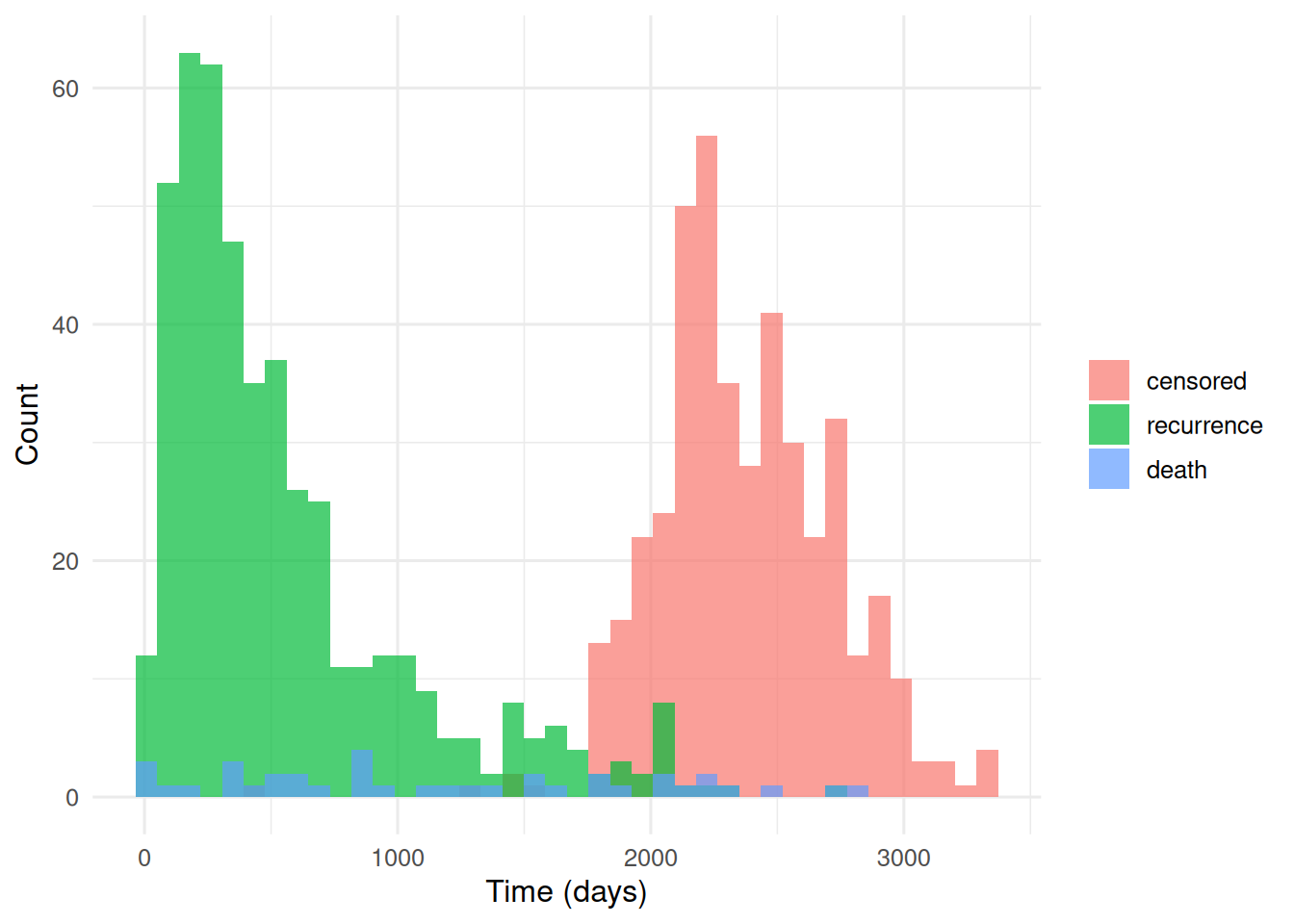
3. Assumptions
Non-informative censoring within cause; proportional hazards for Fine-Gray; for multistate, Markov transitions (future depends only on current state).
4. Conduct
strata time n.risk estimate std.error 95% CI
Obs 500 203 0.006 0.004 0.001, 0.021
Obs 1,000 161 0.019 0.008 0.008, 0.039
Obs 1,500 139 0.029 0.009 0.014, 0.052
Obs 2,000 115 0.032 0.010 0.016, 0.056
Obs 2,500 40 0.044 0.013 0.023, 0.074
Obs 3,000 5 0.075 0.034 0.026, 0.157
Lev 500 199 0.013 0.006 0.004, 0.031
Lev 1,000 157 0.019 0.008 0.008, 0.040
Lev 1,500 145 0.023 0.008 0.010, 0.044
Lev 2,000 121 0.026 0.009 0.012, 0.048
Lev 2,500 49 0.037 0.012 0.018, 0.067
Lev 3,000 4 0.037 0.012 0.018, 0.067
Lev+5FU 500 231 0.016 0.007 0.006, 0.036
Lev+5FU 1,000 198 0.023 0.009 0.010, 0.045
Lev+5FU 1,500 184 0.023 0.009 0.010, 0.045
Lev+5FU 2,000 158 0.037 0.011 0.019, 0.062
Lev+5FU 2,500 62 0.051 0.014 0.029, 0.083
Lev+5FU 3,000 7 0.068 0.022 0.034, 0.118 strata time n.risk estimate std.error 95% CI
Obs 500 203 0.346 0.027 0.294, 0.399
Obs 1,000 161 0.467 0.028 0.411, 0.521
Obs 1,500 139 0.528 0.028 0.471, 0.582
Obs 2,000 115 0.547 0.028 0.491, 0.601
Obs 2,500 40 0.566 0.029 0.508, 0.620
Obs 3,000 5 0.584 0.033 0.517, 0.645
Lev 500 199 0.345 0.027 0.293, 0.398
Lev 1,000 157 0.474 0.028 0.418, 0.529
Lev 1,500 145 0.510 0.028 0.453, 0.564
Lev 2,000 121 0.543 0.028 0.485, 0.596
Lev 2,500 49 0.558 0.029 0.501, 0.612
Lev 3,000 4 0.558 0.029 0.501, 0.612
Lev+5FU 500 231 0.224 0.024 0.179, 0.272
Lev+5FU 1,000 198 0.326 0.027 0.274, 0.379
Lev+5FU 1,500 184 0.362 0.028 0.308, 0.416
Lev+5FU 2,000 158 0.382 0.028 0.327, 0.437
Lev+5FU 2,500 62 0.394 0.028 0.338, 0.449
Lev+5FU 3,000 7 0.394 0.028 0.338, 0.449 outcome statistic df p.value
recurrence 23.7 2.00 <0.001
death 1.09 2.00 0.58 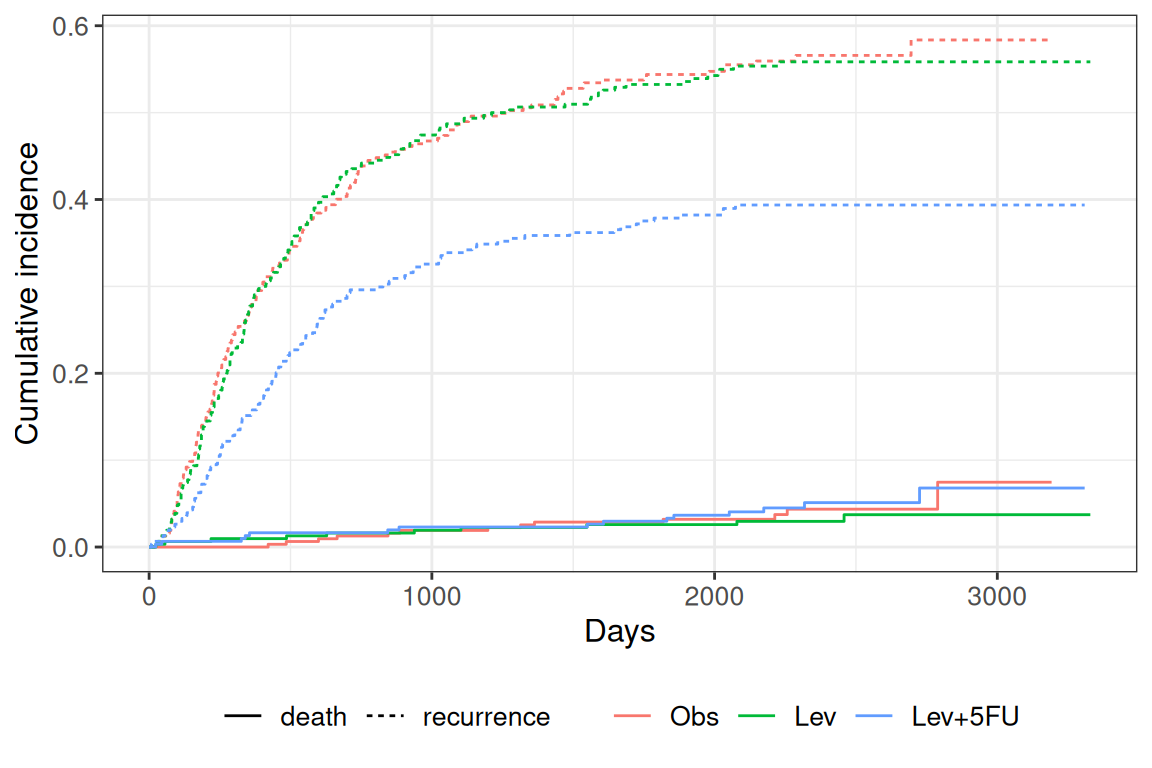
Variable Coef SE HR 95% CI p-value
rxLev -0.014 0.108 0.99 0.80, 1.22 0.90
rxLev+5FU -0.516 0.118 0.60 0.47, 0.75 <0.001
age -0.007 0.004 0.99 0.98, 1.00 0.060
sex -0.113 0.093 0.89 0.75, 1.07 0.22 # Multistate: illness-death using mstate on a simple simulated dataset
n <- 300
sim <- tibble(
id = seq_len(n),
t_ill = rexp(n, rate = 0.02),
t_death = rexp(n, rate = 0.01)
) |>
mutate(
t1 = pmin(t_ill, t_death, 50),
s1 = if_else(t_ill < t_death & t_ill <= 50, 1L, 0L),
s2 = if_else(t_death <= 50 & t_death < t_ill, 1L, 0L)
)
tmat <- transMat(x = list(c(2, 3), c(3), c()),
names = c("healthy", "ill", "dead"))
tmat to
from healthy ill dead
healthy NA 1 2
ill NA NA 3
dead NA NA NA5. Concluding statement
Cause-specific cumulative incidence functions differed by treatment arm in the colon data, with the Fine-Gray subdistribution hazard estimated above. The three-state illness- death transition matrix (healthy → ill → dead) shows how
mstaterepresents the problem; the full fit is straightforward once the long-format dataset is built viamsprep.
Common pitfalls
- Reporting 1 − KM as the cumulative incidence in the presence of competing risks.
- Using Fine-Gray when the question is aetiological.
- Reporting both cause-specific and Fine-Gray hazard ratios without distinguishing them.
- Assuming Markov transitions in a multistate model when history matters.
Further reading
- Fine JP, Gray RJ (1999), A proportional hazards model for the subdistribution of a competing risk.
- Putter H, Fiocco M, Geskus RB (2007), Tutorial in biostatistics: competing risks and multi-state models.
- Austin PC, Fine JP (2017), Practical recommendations for reporting Fine-Gray model analyses.
Session info
R version 4.4.1 (2024-06-14)
Platform: x86_64-pc-linux-gnu
Running under: Ubuntu 24.04.4 LTS
Matrix products: default
BLAS: /usr/lib/x86_64-linux-gnu/openblas-pthread/libblas.so.3
LAPACK: /usr/lib/x86_64-linux-gnu/openblas-pthread/libopenblasp-r0.3.26.so; LAPACK version 3.12.0
locale:
[1] LC_CTYPE=C.UTF-8 LC_NUMERIC=C LC_TIME=C.UTF-8
[4] LC_COLLATE=C.UTF-8 LC_MONETARY=C.UTF-8 LC_MESSAGES=C.UTF-8
[7] LC_PAPER=C.UTF-8 LC_NAME=C LC_ADDRESS=C
[10] LC_TELEPHONE=C LC_MEASUREMENT=C.UTF-8 LC_IDENTIFICATION=C
time zone: UTC
tzcode source: system (glibc)
attached base packages:
[1] stats graphics grDevices utils datasets methods base
other attached packages:
[1] mstate_0.3.3 ggsurvfit_1.2.0 tidycmprsk_1.1.2 survival_3.6-4
[5] lubridate_1.9.5 forcats_1.0.1 stringr_1.6.0 dplyr_1.2.1
[9] purrr_1.2.2 readr_2.2.0 tidyr_1.3.2 tibble_3.3.1
[13] ggplot2_4.0.3 tidyverse_2.0.0
loaded via a namespace (and not attached):
[1] generics_0.1.4 gtsummary_2.5.0 stringi_1.8.7
[4] lattice_0.22-6 hms_1.1.4 digest_0.6.39
[7] magrittr_2.0.5 evaluate_1.0.5 grid_4.4.1
[10] timechange_0.4.0 RColorBrewer_1.1-3 cards_0.7.1
[13] fastmap_1.2.0 jsonlite_2.0.0 Matrix_1.7-0
[16] backports_1.5.1 scales_1.4.0 cli_3.6.6
[19] rlang_1.2.0 hardhat_1.4.3 splines_4.4.1
[22] withr_3.0.2 yaml_2.3.12 otel_0.2.0
[25] tools_4.4.1 tzdb_0.5.0 broom_1.0.12
[28] vctrs_0.7.3 R6_2.6.1 lifecycle_1.0.5
[31] htmlwidgets_1.6.4 pkgconfig_2.0.3 pillar_1.11.1
[34] gtable_0.3.6 glue_1.8.1 data.table_1.18.2.1
[37] xfun_0.57 tidyselect_1.2.1 knitr_1.51
[40] farver_2.1.2 cmprsk_2.2-12 htmltools_0.5.9
[43] labeling_0.4.3 rmarkdown_2.31 compiler_4.4.1
[46] S7_0.2.2