Week 4, Session 1 — Systematic reviews and PRISMA
Course 3 — #courses
Note
Workflow lab: Goal → Approach → Execution → Check → Report.
Learning objectives
- Frame a systematic-review question using PICO.
- Draft a reproducible search strategy across at least two databases.
- Produce a PRISMA flow diagram from counts at each screening stage.
Prerequisites
None beyond reading a paper.
Background
A systematic review is itself a study: it has a protocol, a pre-specified search strategy, explicit inclusion and exclusion criteria, and a plan for synthesis. The PRISMA 2020 statement is the reporting standard, which means the diagram showing what you searched, what you excluded, and why is not optional. PROSPERO (https://www.crd.york.ac.uk/prospero/) is the standard registry for review protocols — registering early protects you from post-hoc drift.
Good search strategies combine controlled-vocabulary terms (MeSH on PubMed, Emtree on Embase) with free-text terms, connect concepts with Boolean AND, and expand synonyms with OR. A librarian-reviewed strategy typically outperforms a DIY one by a wide margin in recall; co-authoring with a health-sciences librarian is the single best investment in review quality.
Setup
1. Goal
Illustrate a PICO-framed question, sketch a search strategy, and simulate screening counts to generate a PRISMA flow diagram.
2. Approach
A fictional question in PICO form:
In adults with type 2 diabetes (P), does metformin (I) compared with placebo (C) reduce all-cause mortality (O) at 24 months?
A sketch of a search strategy (PubMed syntax):
("Diabetes Mellitus, Type 2"[MeSH] OR "type 2 diabetes"[tiab])
AND (metformin[MeSH] OR metformin[tiab])
AND (randomised[tiab] OR randomized[tiab] OR trial[tiab] OR RCT[tiab])3. Execution — simulated screening counts
# A tibble: 7 × 2
stage n
<chr> <dbl>
1 Identified (PubMed) 812
2 Identified (Embase) 640
3 After deduplication 1098
4 Title/abstract screened 1098
5 Full-text assessed 84
6 Included in qualitative synthesis 22
7 Included in meta-analysis 18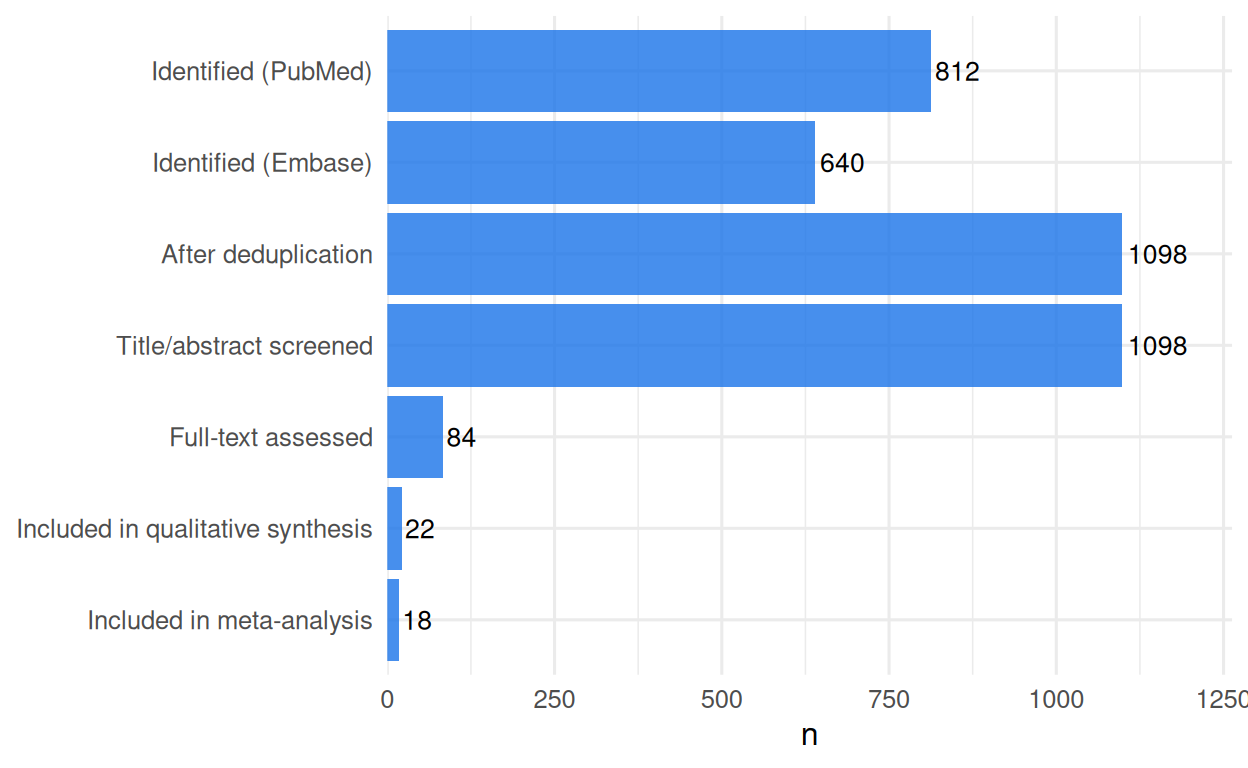
4. Check
PRISMA 2020 checklist pointer (items most often missed):
- Item 5: eligibility criteria pre-specified.
- Item 6: information sources with date of last search.
- Item 7: complete search strategy for every database, including limits.
- Item 16a: reasons for excluding full-text articles.
5. Report
A systematic review of the effect of metformin on all-cause mortality in adults with type 2 diabetes was conducted following PRISMA 2020. Database searches identified 812 records from PubMed and 640 from Embase. After deduplication, 1098 records were screened by title and abstract; 84 full texts were assessed, and 22 studies were included in the qualitative synthesis, of which 18 were pooled by meta-analysis.
Common pitfalls
- Ad-hoc searches that cannot be re-run.
- Failing to double-screen at each stage (reviewer drift).
- Reporting included-study counts without the flow that produced them.
Further reading
- Page MJ et al. (2021). The PRISMA 2020 statement. BMJ.
- Cochrane Handbook for Systematic Reviews of Interventions.
Session info
R version 4.4.1 (2024-06-14)
Platform: x86_64-pc-linux-gnu
Running under: Ubuntu 24.04.4 LTS
Matrix products: default
BLAS: /usr/lib/x86_64-linux-gnu/openblas-pthread/libblas.so.3
LAPACK: /usr/lib/x86_64-linux-gnu/openblas-pthread/libopenblasp-r0.3.26.so; LAPACK version 3.12.0
locale:
[1] LC_CTYPE=C.UTF-8 LC_NUMERIC=C LC_TIME=C.UTF-8
[4] LC_COLLATE=C.UTF-8 LC_MONETARY=C.UTF-8 LC_MESSAGES=C.UTF-8
[7] LC_PAPER=C.UTF-8 LC_NAME=C LC_ADDRESS=C
[10] LC_TELEPHONE=C LC_MEASUREMENT=C.UTF-8 LC_IDENTIFICATION=C
time zone: UTC
tzcode source: system (glibc)
attached base packages:
[1] stats graphics grDevices utils datasets methods base
other attached packages:
[1] lubridate_1.9.5 forcats_1.0.1 stringr_1.6.0 dplyr_1.2.1
[5] purrr_1.2.2 readr_2.2.0 tidyr_1.3.2 tibble_3.3.1
[9] ggplot2_4.0.3 tidyverse_2.0.0
loaded via a namespace (and not attached):
[1] gtable_0.3.6 jsonlite_2.0.0 compiler_4.4.1 tidyselect_1.2.1
[5] scales_1.4.0 yaml_2.3.12 fastmap_1.2.0 R6_2.6.1
[9] labeling_0.4.3 generics_0.1.4 knitr_1.51 htmlwidgets_1.6.4
[13] pillar_1.11.1 RColorBrewer_1.1-3 tzdb_0.5.0 rlang_1.2.0
[17] utf8_1.2.6 stringi_1.8.7 xfun_0.57 S7_0.2.2
[21] otel_0.2.0 timechange_0.4.0 cli_3.6.6 withr_3.0.2
[25] magrittr_2.0.5 digest_0.6.39 grid_4.4.1 hms_1.1.4
[29] lifecycle_1.0.5 vctrs_0.7.3 evaluate_1.0.5 glue_1.8.1
[33] farver_2.1.2 rmarkdown_2.31 tools_4.4.1 pkgconfig_2.0.3
[37] htmltools_0.5.9