Week 4, Session 3 — Network meta-analysis
Course 3 — #courses
Note
Inference lab: Hypothesis → Visualise → Assumptions → Conduct → Conclude.
Learning objectives
- Fit a network meta-analysis (NMA) using
netmeta. - Draw and read the network graph.
- Derive SUCRA rankings and interpret them cautiously.
Prerequisites
Pairwise meta-analysis (W4 S2).
Background
Network meta-analysis extends pairwise meta-analysis to more than two treatments by combining direct comparisons (from head-to-head trials) with indirect comparisons (inferred through a shared comparator). The central assumption is transitivity: the studies compared indirectly are exchangeable — no moderator distinguishes them in a way that systematically shifts the effect. The statistical analogue, consistency, checks whether direct and indirect estimates agree.
SUCRA (surface under the cumulative ranking curve) summarises each treatment’s rank distribution on a 0-1 scale, where 1 is best. SUCRA values compress a lot of uncertainty into a single number; they are useful for signalling but should be reported with interval estimates for every pairwise comparison.
Setup
1. Hypothesis
H₀: all treatments in the network have equal efficacy. H₁: at least two differ.
2. Visualise
TE seTE treat1.long treat2.long treat1 treat2 studlab
1 -1.90 0.1414 Metformin Placebo metf plac DeFronzo1995
2 -0.82 0.0992 Metformin Placebo metf plac Lewin2007
3 -0.20 0.3579 Metformin Acarbose metf acar Willms1999
4 -1.34 0.1435 Rosiglitazone Placebo rosi plac Davidson2007
5 -1.10 0.1141 Rosiglitazone Placebo rosi plac Wolffenbuttel1999
6 -1.30 0.1268 Pioglitazone Placebo piog plac Kipnes2001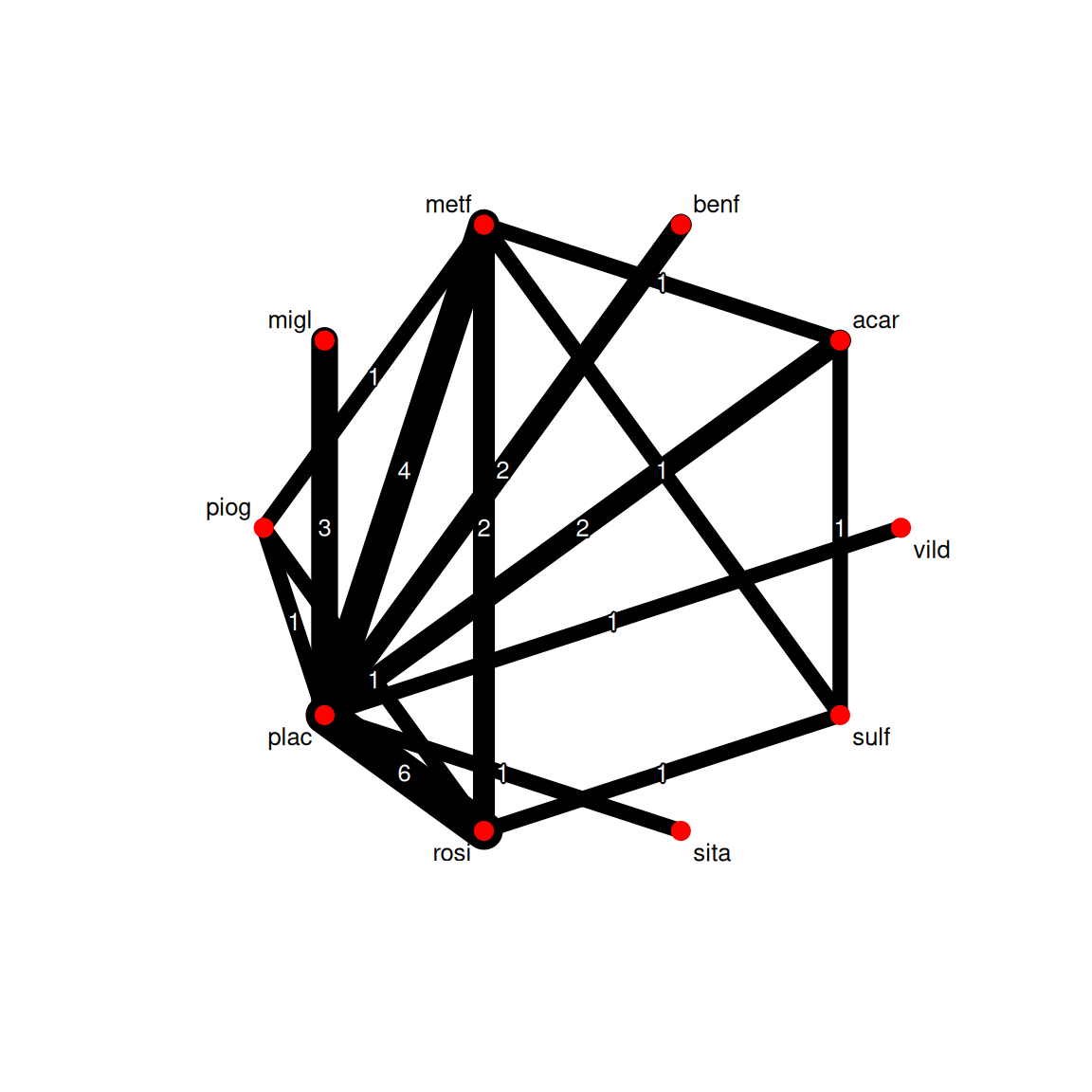
3. Assumptions
Q statistics to assess homogeneity / consistency
Q df p-value
Total 96.99 18 < 0.0001
Within designs 74.46 11 < 0.0001
Between designs 22.53 7 0.0021
Design-specific decomposition of within-designs Q statistic
Design Q df p-value
plac:metf 42.16 2 < 0.0001
plac:rosi 21.27 5 0.0007
plac:benf 4.38 1 0.0363
plac:migl 6.45 2 0.0398
metf:rosi 0.19 1 0.6655
Between-designs Q statistic after detaching of single designs
(influential designs have p-value markedly different from 0.0021)
Detached design Q df p-value
rosi:sulf 6.77 6 0.3425
metf:sulf 7.51 6 0.2760
plac:rosi 16.29 6 0.0123
metf:piog 17.13 6 0.0088
plac:piog 17.25 6 0.0084
plac:metf 22.07 6 0.0012
plac:acar 22.44 6 0.0010
piog:rosi 22.48 6 0.0010
acar:sulf 22.52 6 0.0010
metf:rosi 22.52 6 0.0010
plac:acar:metf 22.38 5 0.0004
Q statistic to assess consistency under the assumption of
a full design-by-treatment interaction random effects model
Q df p-value tau.within tau2.within
Between designs 2.19 7 0.9483 0.3797 0.1442The within-design heterogeneity and between-design inconsistency variances are the key diagnostics.
4. Conduct
Original data (with adjusted standard errors for multi-arm studies):
treat1 treat2 TE seTE seTE.adj narms multiarm
DeFronzo1995 metf plac -1.9000 0.1414 0.3588 2
Lewin2007 metf plac -0.8200 0.0992 0.3443 2
Willms1999 acar metf 0.2000 0.3579 0.5574 3 *
Davidson2007 plac rosi 1.3400 0.1435 0.3596 2
Wolffenbuttel1999 plac rosi 1.1000 0.1141 0.3489 2
Kipnes2001 piog plac -1.3000 0.1268 0.3533 2
Kerenyi2004 plac rosi 0.7700 0.1078 0.3469 2
Hanefeld2004 metf piog -0.1600 0.0849 0.3405 2
Derosa2004 piog rosi 0.1000 0.1831 0.3772 2
Baksi2004 plac rosi 1.3000 0.1014 0.3450 2
Rosenstock2008 plac rosi 1.0900 0.2263 0.3999 2
Zhu2003 plac rosi 1.5000 0.1624 0.3675 2
Yang2003 metf rosi 0.1400 0.2239 0.3986 2
Vongthavaravat2002 rosi sulf -1.2000 0.1436 0.3596 2
Oyama2008 acar sulf -0.4000 0.1549 0.3643 2
Costa1997 acar plac -0.8000 0.1432 0.3595 2
Hermansen2007 plac sita 0.5700 0.1291 0.3541 2
Garber2008 plac vild 0.7000 0.1273 0.3534 2
Alex1998 metf sulf -0.3700 0.1184 0.3503 2
Johnston1994 migl plac -0.7400 0.1839 0.3775 2
Johnston1998a migl plac -1.4100 0.2235 0.3983 2
Kim2007 metf rosi -0.0000 0.2339 0.4043 2
Johnston1998b migl plac -0.6800 0.2828 0.4344 2
Gonzalez-Ortiz2004 metf plac -0.4000 0.4356 0.5463 2
Stucci1996 benf plac -0.2300 0.3467 0.4785 2
Moulin2006 benf plac -1.0100 0.1366 0.3569 2
Willms1999 metf plac -1.2000 0.3758 0.5802 3 *
Willms1999 acar plac -1.0000 0.4669 0.8122 3 *
Number of treatment arms per study (by decreasing number of arms):
narms multiarm
Willms1999 3 *
DeFronzo1995 2
Lewin2007 2
Davidson2007 2
Wolffenbuttel1999 2
Kipnes2001 2
Kerenyi2004 2
Hanefeld2004 2
Derosa2004 2
Baksi2004 2
Rosenstock2008 2
Zhu2003 2
Yang2003 2
Vongthavaravat2002 2
Oyama2008 2
Costa1997 2
Hermansen2007 2
Garber2008 2
Alex1998 2
Johnston1994 2
Johnston1998a 2
Kim2007 2
Johnston1998b 2
Gonzalez-Ortiz2004 2
Stucci1996 2
Moulin2006 2
Results (random effects model):
treat1 treat2 MD 95%-CI
DeFronzo1995 metf plac -1.1268 [-1.4291; -0.8244]
Lewin2007 metf plac -1.1268 [-1.4291; -0.8244]
Willms1999 acar metf 0.2850 [-0.2208; 0.7908]
Davidson2007 plac rosi 1.2335 [ 0.9830; 1.4839]
Wolffenbuttel1999 plac rosi 1.2335 [ 0.9830; 1.4839]
Kipnes2001 piog plac -1.1291 [-1.5596; -0.6986]
Kerenyi2004 plac rosi 1.2335 [ 0.9830; 1.4839]
Hanefeld2004 metf piog 0.0023 [-0.4398; 0.4444]
Derosa2004 piog rosi 0.1044 [-0.3347; 0.5435]
Baksi2004 plac rosi 1.2335 [ 0.9830; 1.4839]
Rosenstock2008 plac rosi 1.2335 [ 0.9830; 1.4839]
Zhu2003 plac rosi 1.2335 [ 0.9830; 1.4839]
Yang2003 metf rosi 0.1067 [-0.2170; 0.4304]
Vongthavaravat2002 rosi sulf -0.8169 [-1.2817; -0.3521]
Oyama2008 acar sulf -0.4252 [-0.9456; 0.0951]
Costa1997 acar plac -0.8418 [-1.3236; -0.3600]
Hermansen2007 plac sita 0.5700 [-0.1240; 1.2640]
Garber2008 plac vild 0.7000 [ 0.0073; 1.3927]
Alex1998 metf sulf -0.7102 [-1.1713; -0.2491]
Johnston1994 migl plac -0.9497 [-1.4040; -0.4955]
Johnston1998a migl plac -0.9497 [-1.4040; -0.4955]
Kim2007 metf rosi 0.1067 [-0.2170; 0.4304]
Johnston1998b migl plac -0.9497 [-1.4040; -0.4955]
Gonzalez-Ortiz2004 metf plac -1.1268 [-1.4291; -0.8244]
Stucci1996 benf plac -0.7311 [-1.2918; -0.1705]
Moulin2006 benf plac -0.7311 [-1.2918; -0.1705]
Willms1999 metf plac -1.1268 [-1.4291; -0.8244]
Willms1999 acar plac -0.8418 [-1.3236; -0.3600]
Number of studies: k = 26
Number of pairwise comparisons: m = 28
Number of treatments: n = 10
Number of designs: d = 15
Random effects model
Treatment estimate (other treatments vs 'plac'):
MD 95%-CI z p-value
acar -0.8418 [-1.3236; -0.3600] -3.42 0.0006
benf -0.7311 [-1.2918; -0.1705] -2.56 0.0106
metf -1.1268 [-1.4291; -0.8244] -7.30 < 0.0001
migl -0.9497 [-1.4040; -0.4955] -4.10 < 0.0001
piog -1.1291 [-1.5596; -0.6986] -5.14 < 0.0001
plac . . . .
rosi -1.2335 [-1.4839; -0.9830] -9.65 < 0.0001
sita -0.5700 [-1.2640; 0.1240] -1.61 0.1075
sulf -0.4166 [-0.8887; 0.0556] -1.73 0.0838
vild -0.7000 [-1.3927; -0.0073] -1.98 0.0476
Quantifying heterogeneity / inconsistency:
tau^2 = 0.1087; tau = 0.3297; I^2 = 81.4% [72.0%; 87.7%]
Tests of heterogeneity (within designs) and inconsistency (between designs):
Q d.f. p-value
Total 96.99 18 < 0.0001
Within designs 74.46 11 < 0.0001
Between designs 22.53 7 0.0021
Details of network meta-analysis methods:
- Frequentist graph-theoretical approach
- DerSimonian-Laird estimator for tau^2
- Calculation of I^2 based on Q P-score
rosi 0.8934
metf 0.7818
piog 0.7746
migl 0.6137
acar 0.5203
benf 0.4358
vild 0.4232
sita 0.3331
sulf 0.2103
plac 0.01395. Concluding statement
In a network meta-analysis of 10 treatments across 26 studies (28 pairwise comparisons), the treatment with the highest SUCRA for HbA1c lowering was rosi (SUCRA = 0.89). SUCRA rankings should be interpreted alongside the pairwise effect estimates and their intervals, not as a standalone conclusion.
Common pitfalls
- Treating SUCRA as if it were the effect size.
- Ignoring the transitivity assumption because the output compiled.
- Mixing direct and indirect evidence without quantifying inconsistency.
Further reading
- Rücker G, Schwarzer G. (2015). Ranking treatments in frequentist network meta-analysis.
- Salanti G. (2012). Indirect and mixed-treatment comparison…
Session info
R version 4.4.1 (2024-06-14)
Platform: x86_64-pc-linux-gnu
Running under: Ubuntu 24.04.4 LTS
Matrix products: default
BLAS: /usr/lib/x86_64-linux-gnu/openblas-pthread/libblas.so.3
LAPACK: /usr/lib/x86_64-linux-gnu/openblas-pthread/libopenblasp-r0.3.26.so; LAPACK version 3.12.0
locale:
[1] LC_CTYPE=C.UTF-8 LC_NUMERIC=C LC_TIME=C.UTF-8
[4] LC_COLLATE=C.UTF-8 LC_MONETARY=C.UTF-8 LC_MESSAGES=C.UTF-8
[7] LC_PAPER=C.UTF-8 LC_NAME=C LC_ADDRESS=C
[10] LC_TELEPHONE=C LC_MEASUREMENT=C.UTF-8 LC_IDENTIFICATION=C
time zone: UTC
tzcode source: system (glibc)
attached base packages:
[1] stats graphics grDevices utils datasets methods base
other attached packages:
[1] netmeta_3.4-0 meta_8.3-0 metadat_1.6-0 metabook_0.2-0
[5] lubridate_1.9.5 forcats_1.0.1 stringr_1.6.0 dplyr_1.2.1
[9] purrr_1.2.2 readr_2.2.0 tidyr_1.3.2 tibble_3.3.1
[13] ggplot2_4.0.3 tidyverse_2.0.0
loaded via a namespace (and not attached):
[1] gtable_0.3.6 xfun_0.57 htmlwidgets_1.6.4
[4] CompQuadForm_1.4.4 lattice_0.22-6 mathjaxr_2.0-0
[7] tzdb_0.5.0 numDeriv_2016.8-1.1 vctrs_0.7.3
[10] tools_4.4.1 Rdpack_2.6.6 generics_0.1.4
[13] pkgconfig_2.0.3 Matrix_1.7-0 RColorBrewer_1.1-3
[16] S7_0.2.2 lifecycle_1.0.5 compiler_4.4.1
[19] farver_2.1.2 htmltools_0.5.9 yaml_2.3.12
[22] pillar_1.11.1 nloptr_2.2.1 MASS_7.3-60.2
[25] reformulas_0.4.4 abind_1.4-8 boot_1.3-30
[28] nlme_3.1-164 tidyselect_1.2.1 digest_0.6.39
[31] mvtnorm_1.3-7 stringi_1.8.7 splines_4.4.1
[34] magic_1.6-1 fastmap_1.2.0 grid_4.4.1
[37] colorspace_2.1-2 cli_3.6.6 metafor_5.0-1
[40] magrittr_2.0.5 withr_3.0.2 scales_1.4.0
[43] timechange_0.4.0 rmarkdown_2.31 igraph_2.3.1
[46] otel_0.2.0 lme4_2.0-1 hms_1.1.4
[49] evaluate_1.0.5 knitr_1.51 rbibutils_2.4.1
[52] rlang_1.2.0 Rcpp_1.1.1-1.1 glue_1.8.1
[55] xml2_1.5.2 minqa_1.2.8 jsonlite_2.0.0
[58] R6_2.6.1