Week 1, Session 4 — Clustering: k-means, hierarchical, model-based
Course 4 — #courses
Note
Workflow labs use the variant template: Goal → Approach → Execution → Check → Report.
Learning objectives
- Fit k-means and hierarchical clusterings, and choose k by elbow and silhouette.
- Fit a model-based clustering with
mclustand select the number of components by BIC. - Explain why “number of clusters” is often not the right question.
Prerequisites
PCA from Session 3; distance and linkage from Course 1.
Background
Clustering is a search for structure in unlabelled data, and unlike classification it has no gold standard. Any algorithm will return a partition if asked to, and the partitions it returns depend on assumptions about what a cluster is. K-means assumes roughly spherical, equal-variance clusters and needs k in advance. Hierarchical clustering produces a nested sequence of partitions at every k, which you cut at a level you choose. Model-based clustering fits a Gaussian mixture and selects k by information criterion, which at least makes the assumption explicit.
In biomedicine, the honest version of “how many clusters?” is usually “is there more than one?” A good clustering lab focuses on stability and interpretability of the partition rather than on the number returned.
When you report a clustering, report the number of clusters, the criterion used to select it, a visualisation in a 2-D projection (PCA or UMAP), and the size and key features of each cluster. Do not report p-values for cluster labels against the data that produced the labels; that circularity is a classic error.
Setup
1. Goal
Cluster a simulated dataset with three known groups using three algorithms, and see which recovers the structure.
2. Approach
Three Gaussian blobs in 5 dimensions with different sizes.
make_blob <- function(n, mu, sd = 1) matrix(rnorm(n * 5, 0, sd), n, 5) +
rep(mu, each = n)
X <- rbind(
make_blob(60, c(0, 0, 0, 0, 0)),
make_blob(40, c(3, 0, 0, 0, 0)),
make_blob(30, c(0, 3, 0, 0, 0))
)
true_lbl <- rep(c("A", "B", "C"), c(60, 40, 30))
pc <- prcomp(X)$x[, 1:2]
tibble(PC1 = pc[, 1], PC2 = pc[, 2], group = true_lbl) |>
ggplot(aes(PC1, PC2, colour = group)) + geom_point(alpha = 0.7)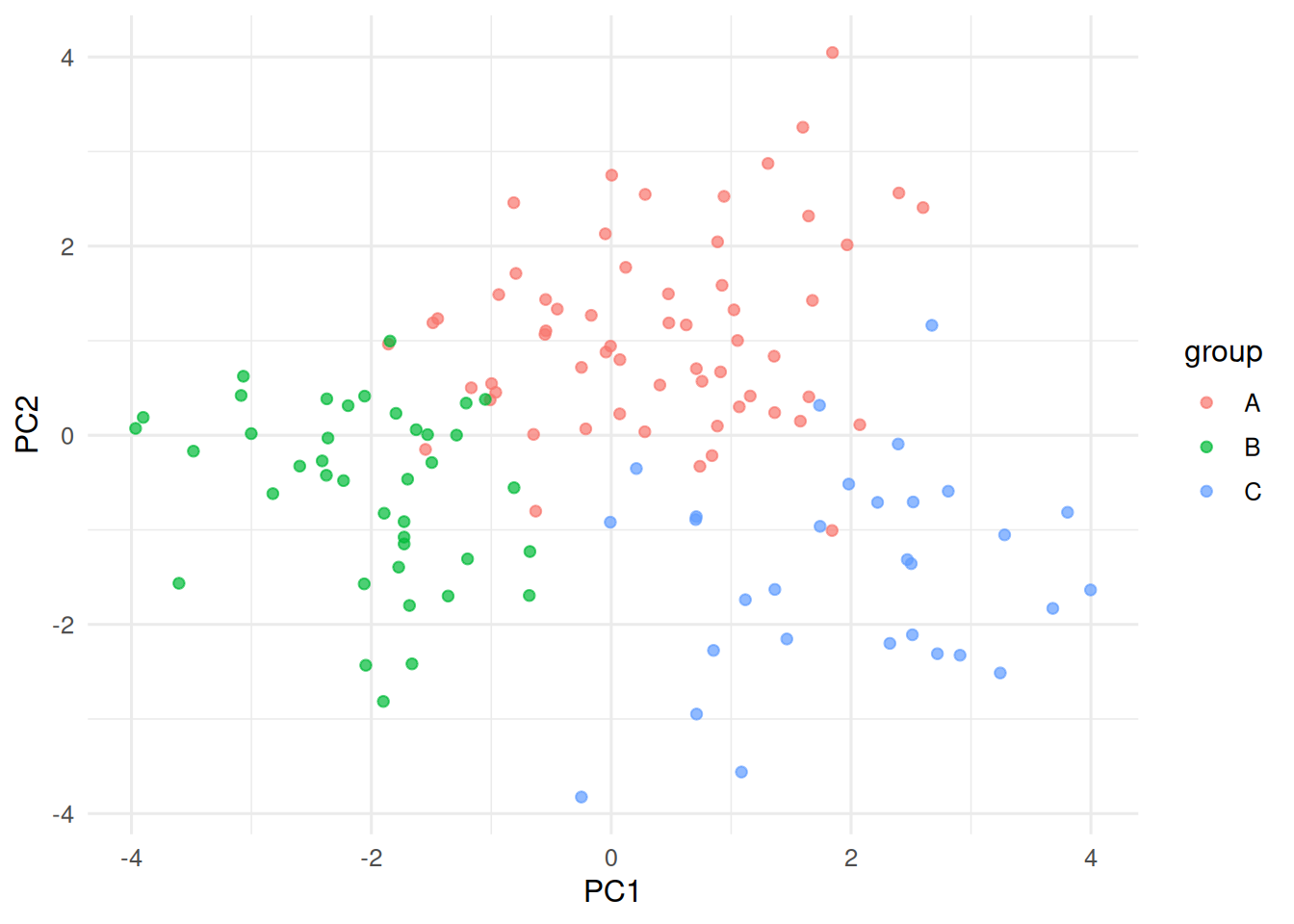
3. Execution
K-means with k sweep; silhouette.
wss <- sapply(2:8, function(k) kmeans(X, centers = k, nstart = 10)$tot.withinss)
sil <- sapply(2:8, function(k) {
km <- kmeans(X, centers = k, nstart = 10)
mean(silhouette(km$cluster, dist(X))[, 3])
})
tibble(k = 2:8, wss = wss, sil = sil) |>
pivot_longer(-k) |>
ggplot(aes(k, value)) + geom_line() + geom_point() +
facet_wrap(~ name, scales = "free_y")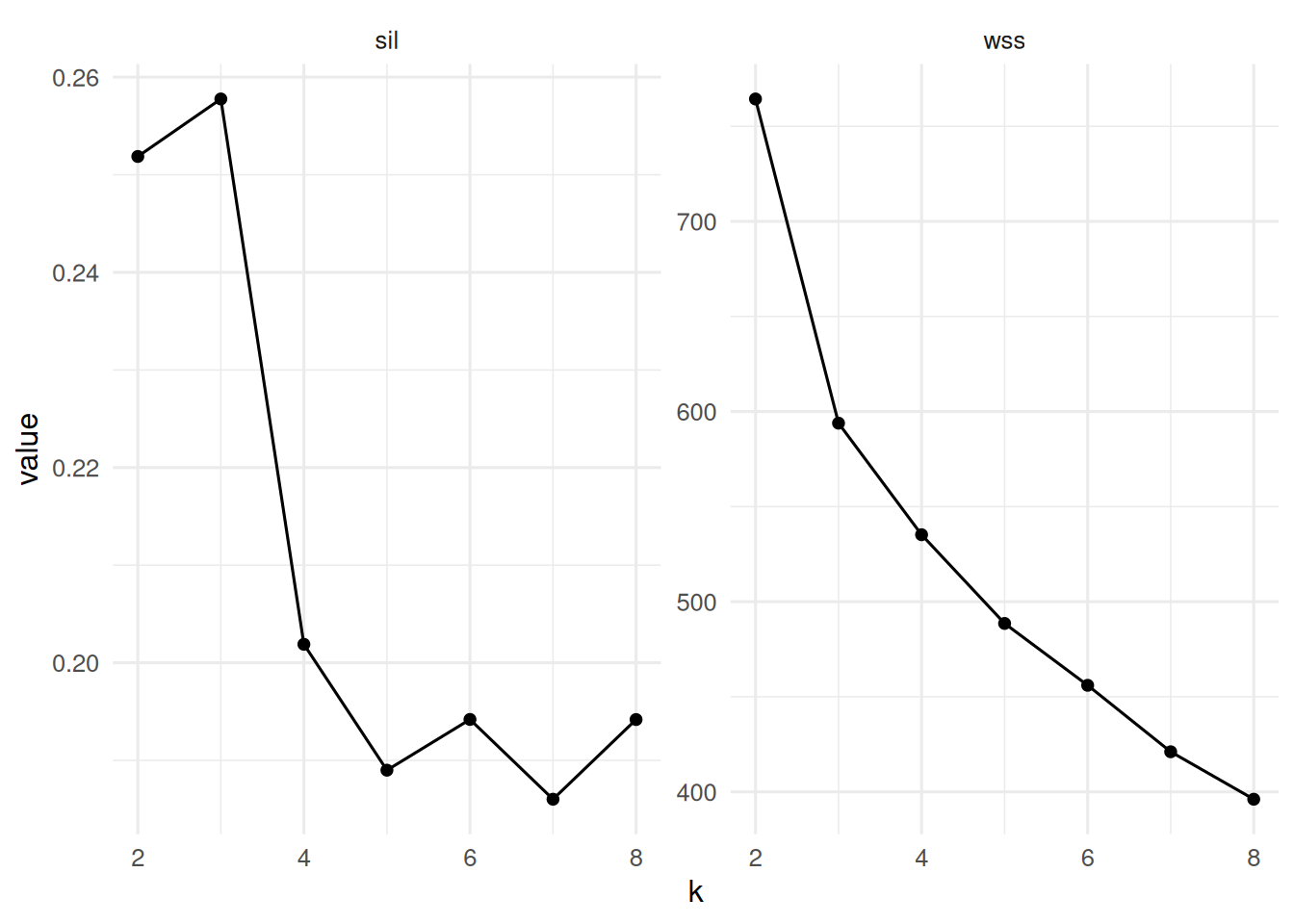
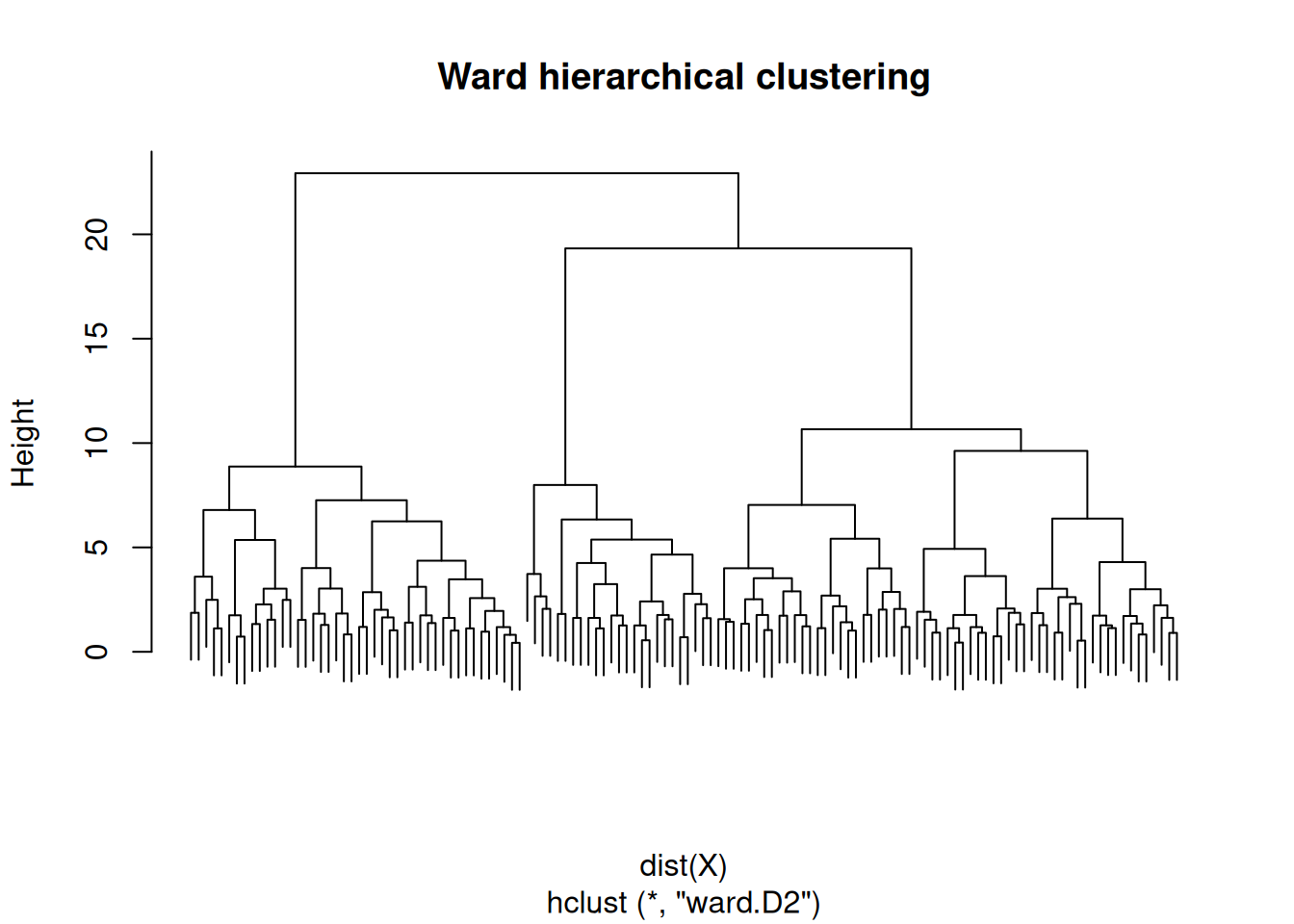
Hierarchical with Ward linkage.
4. Check
Model-based clustering by BIC.
----------------------------------------------------
Gaussian finite mixture model fitted by EM algorithm
----------------------------------------------------
Mclust EII (spherical, equal volume) model with 3 components:
log-likelihood n df BIC ICL
-1018.943 130 18 -2125.503 -2151.66
Clustering table:
1 2 3
51 53 26 5. Report
On three simulated Gaussian blobs (n = 130 in R⁵), model-based clustering selected G = 3 by BIC; k-means with k = 3 maximised mean silhouette (0.26). Hierarchical clustering with Ward linkage produced a dendrogram consistent with three primary branches.
When the generating model is actually Gaussian, mclust tends to win. On real data, no single method is best; the test is whether the partition is stable under resampling and interpretable in terms of the features.
Common pitfalls
- Treating k-means output as ground truth; it is an optimisation answer given k.
- Running hierarchical clustering on a very large dataset without noting the O(n²) memory cost.
- Testing post-hoc differences between clusters with the same data that produced them.
Further reading
- Fraley C, Raftery AE (2002), Model-based clustering, discriminant analysis, and density estimation.
- Hastie, Tibshirani, Friedman, ESL, §14.3.
Session info
R version 4.4.1 (2024-06-14)
Platform: x86_64-pc-linux-gnu
Running under: Ubuntu 24.04.4 LTS
Matrix products: default
BLAS: /usr/lib/x86_64-linux-gnu/openblas-pthread/libblas.so.3
LAPACK: /usr/lib/x86_64-linux-gnu/openblas-pthread/libopenblasp-r0.3.26.so; LAPACK version 3.12.0
locale:
[1] LC_CTYPE=C.UTF-8 LC_NUMERIC=C LC_TIME=C.UTF-8
[4] LC_COLLATE=C.UTF-8 LC_MONETARY=C.UTF-8 LC_MESSAGES=C.UTF-8
[7] LC_PAPER=C.UTF-8 LC_NAME=C LC_ADDRESS=C
[10] LC_TELEPHONE=C LC_MEASUREMENT=C.UTF-8 LC_IDENTIFICATION=C
time zone: UTC
tzcode source: system (glibc)
attached base packages:
[1] stats graphics grDevices utils datasets methods base
other attached packages:
[1] cluster_2.1.8.2 mclust_6.1.2 lubridate_1.9.5 forcats_1.0.1
[5] stringr_1.6.0 dplyr_1.2.1 purrr_1.2.2 readr_2.2.0
[9] tidyr_1.3.2 tibble_3.3.1 ggplot2_4.0.3 tidyverse_2.0.0
loaded via a namespace (and not attached):
[1] gtable_0.3.6 jsonlite_2.0.0 compiler_4.4.1 tidyselect_1.2.1
[5] scales_1.4.0 yaml_2.3.12 fastmap_1.2.0 R6_2.6.1
[9] labeling_0.4.3 generics_0.1.4 knitr_1.51 htmlwidgets_1.6.4
[13] pillar_1.11.1 RColorBrewer_1.1-3 tzdb_0.5.0 rlang_1.2.0
[17] stringi_1.8.7 xfun_0.57 S7_0.2.2 otel_0.2.0
[21] timechange_0.4.0 cli_3.6.6 withr_3.0.2 magrittr_2.0.5
[25] digest_0.6.39 grid_4.4.1 hms_1.1.4 lifecycle_1.0.5
[29] vctrs_0.7.3 evaluate_1.0.5 glue_1.8.1 farver_2.1.2
[33] rmarkdown_2.31 tools_4.4.1 pkgconfig_2.0.3 htmltools_0.5.9