Week 2, Session 1 — Trees, random forests, gradient boosting
Course 4 — #courses
Note
Workflow labs use the variant template: Goal → Approach → Execution → Check → Report.
Learning objectives
- Fit a single decision tree, a random forest, and a gradient-boosted model on the same data, and compare their error.
- Read variable-importance output critically and know its biases.
- Explain when a tree ensemble is preferable to a linear model and when it is not.
Prerequisites
CV from Session 1; regularisation from Session 2.
Background
Tree-based models divide feature space by axis-aligned splits and fit a constant within each region. A single tree is interpretable but high variance; averaging many trees with resampling and random feature subsets gives random forests, which are among the strongest off-the-shelf methods on tabular data. Gradient boosting takes a different route: it fits trees sequentially, each correcting the residuals of the last, and usually wins on tabular benchmarks when carefully tuned.
Tree methods make few assumptions about the functional form of the response and handle interactions automatically. Their limitations are extrapolation (flat beyond the training range), variable-importance measures that can be misleading when features are correlated, and a general tendency to look better than they generalise unless honest resampling is used.
When reporting a random forest or gradient-boosted model, report the tuning procedure, the out-of-bag or CV error, the variable-importance method, and — if the goal is scientific — a permutation-based importance to cross-check the default Gini/gain importance.
Setup
1. Goal
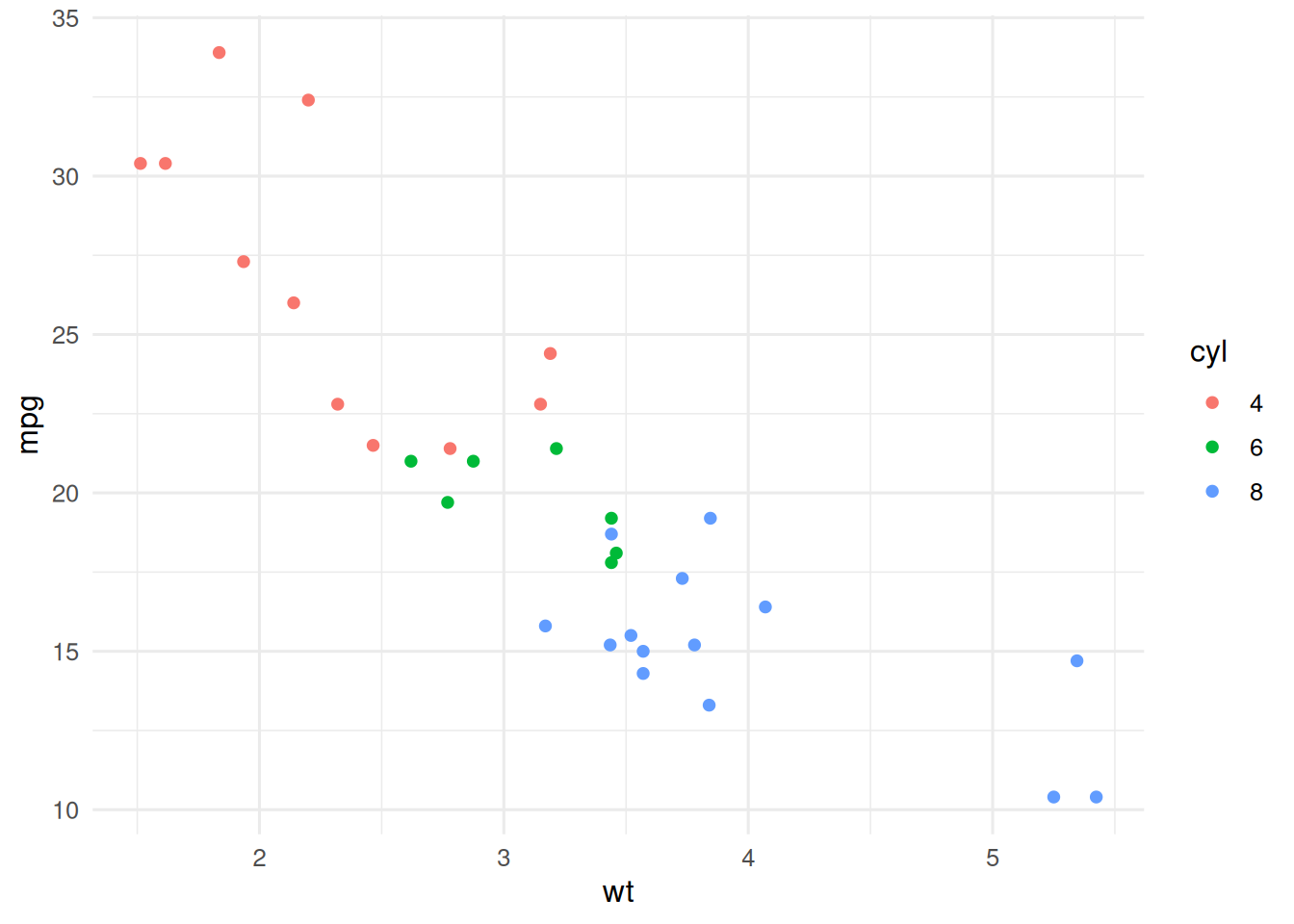
Predict mpg from mtcars with a single tree, a random forest, and gradient boosting; compare errors and importances.
2. Approach
3. Execution
4. Check
Honest error: CV for the tree and xgboost, OOB for ranger.
K <- 5
folds <- sample(rep(1:K, length.out = nrow(d)))
mse <- function(y, yhat) mean((y - yhat)^2)
get_err <- function() {
err_tree <- err_rf <- err_xg <- numeric(K)
for (k in 1:K) {
tr <- folds != k; te <- folds == k
f_tr <- d[tr, ]; f_te <- d[te, ]
t1 <- rpart(mpg ~ ., data = f_tr, cp = 0.01)
r1 <- ranger(mpg ~ ., data = f_tr, num.trees = 500)
x1 <- xgboost(
data = as.matrix(f_tr[, -1]), label = f_tr$mpg,
nrounds = 100, eta = 0.05, max_depth = 3,
objective = "reg:squarederror", verbose = 0
)
err_tree[k] <- mse(f_te$mpg, predict(t1, f_te))
err_rf[k] <- mse(f_te$mpg, predict(r1, f_te)$predictions)
err_xg[k] <- mse(f_te$mpg, predict(x1, as.matrix(f_te[, -1])))
}
c(tree = mean(err_tree), rf = mean(err_rf), xgb = mean(err_xg))
}
errs <- get_err()
errs tree rf xgb
13.900260 4.814310 7.995655 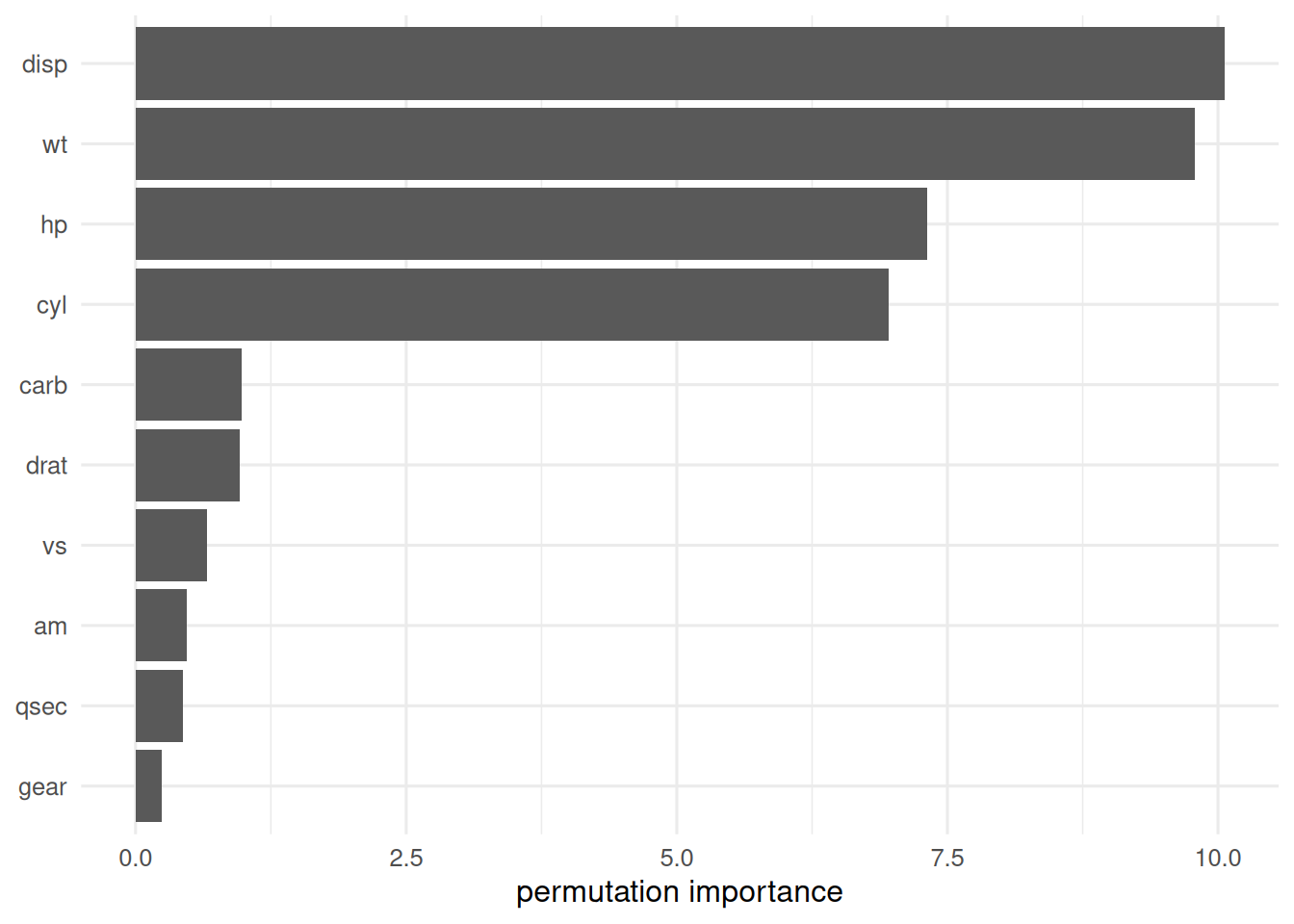
5. Report
On
mtcars(n = 32), 5-fold CV MSEs were 13.9 (single tree), 4.81 (random forest), and 8 (gradient boosting). Permutation importance from the random forest identified weight and displacement as the dominant predictors.
The CV errors are close because the sample is tiny and the problem is close to linear. On larger tabular problems the gap opens and gradient boosting typically wins after tuning; but the tiny-sample regime is where trees and boosting most often look like magic and almost always lie.
Common pitfalls
- Reporting the training error of a forest; always use OOB or CV.
- Trusting Gini/gain importance for correlated predictors.
- Letting xgboost overfit with low
etaand highnroundswithout early stopping.
Further reading
- Breiman L (2001), Random forests.
- Chen T, Guestrin C (2016), XGBoost: A scalable tree boosting system.
Session info
R version 4.4.1 (2024-06-14)
Platform: x86_64-pc-linux-gnu
Running under: Ubuntu 24.04.4 LTS
Matrix products: default
BLAS: /usr/lib/x86_64-linux-gnu/openblas-pthread/libblas.so.3
LAPACK: /usr/lib/x86_64-linux-gnu/openblas-pthread/libopenblasp-r0.3.26.so; LAPACK version 3.12.0
locale:
[1] LC_CTYPE=C.UTF-8 LC_NUMERIC=C LC_TIME=C.UTF-8
[4] LC_COLLATE=C.UTF-8 LC_MONETARY=C.UTF-8 LC_MESSAGES=C.UTF-8
[7] LC_PAPER=C.UTF-8 LC_NAME=C LC_ADDRESS=C
[10] LC_TELEPHONE=C LC_MEASUREMENT=C.UTF-8 LC_IDENTIFICATION=C
time zone: UTC
tzcode source: system (glibc)
attached base packages:
[1] stats graphics grDevices utils datasets methods base
other attached packages:
[1] xgboost_3.2.1.1 ranger_0.18.0 rpart_4.1.23 lubridate_1.9.5
[5] forcats_1.0.1 stringr_1.6.0 dplyr_1.2.1 purrr_1.2.2
[9] readr_2.2.0 tidyr_1.3.2 tibble_3.3.1 ggplot2_4.0.3
[13] tidyverse_2.0.0
loaded via a namespace (and not attached):
[1] Matrix_1.7-0 gtable_0.3.6 jsonlite_2.0.0
[4] compiler_4.4.1 Rcpp_1.1.1-1.1 tidyselect_1.2.1
[7] parallel_4.4.1 scales_1.4.0 yaml_2.3.12
[10] fastmap_1.2.0 lattice_0.22-6 R6_2.6.1
[13] labeling_0.4.3 generics_0.1.4 knitr_1.51
[16] htmlwidgets_1.6.4 pillar_1.11.1 RColorBrewer_1.1-3
[19] tzdb_0.5.0 rlang_1.2.0 stringi_1.8.7
[22] xfun_0.57 S7_0.2.2 otel_0.2.0
[25] timechange_0.4.0 cli_3.6.6 withr_3.0.2
[28] magrittr_2.0.5 digest_0.6.39 grid_4.4.1
[31] hms_1.1.4 lifecycle_1.0.5 vctrs_0.7.3
[34] data.table_1.18.2.1 evaluate_1.0.5 glue_1.8.1
[37] farver_2.1.2 rmarkdown_2.31 tools_4.4.1
[40] pkgconfig_2.0.3 htmltools_0.5.9