Week 3, Session 4 — Survival ML (random survival forest, DeepSurv)
Course 4 — #courses
Note
Workflow labs use the variant template: Goal → Approach → Execution → Check → Report.
Learning objectives
- Fit a random survival forest and interpret its variable importance and risk stratification.
- Describe the DeepSurv architecture conceptually and when it is preferred over the forest.
- Compare against a Cox model on the same features.
Prerequisites
Cox regression and censoring from Course 2.
Background
Survival analysis deals with time-to-event data in the presence of censoring. The Cox proportional-hazards model is the workhorse linear method; random survival forests (RSF) extend the idea of random forests to right-censored outcomes, using a log-rank split rule and Harrell’s concordance as the default fit criterion. They handle nonlinearities and interactions automatically and are a good first nonparametric benchmark.
DeepSurv is a neural-network generalisation of the Cox partial likelihood: the linear predictor is replaced by a feed-forward network. In small biomedical datasets it rarely beats a well-tuned RSF or a Cox model with splines, but the approach scales to large cohorts and to inputs where a representation must be learned.
Evaluation of survival models uses time-integrated concordance (Harrell’s C), the integrated Brier score, and — increasingly — decision curves at a clinically relevant horizon. Reporting only C on a single test set is weak evidence.
Setup
1. Goal
Fit an RSF on survival::pbc and compare to a Cox model using Harrell’s concordance.
2. Approach
d <- survival::pbc |>
as_tibble() |>
drop_na(bili, albumin, age, edema, protime, stage) |>
mutate(status = ifelse(status == 2, 1, 0))
ggplot(d, aes(time / 365.25, fill = factor(status))) +
geom_histogram(bins = 30, alpha = 0.7, position = "identity") +
labs(x = "follow-up (years)", y = "count", fill = "event")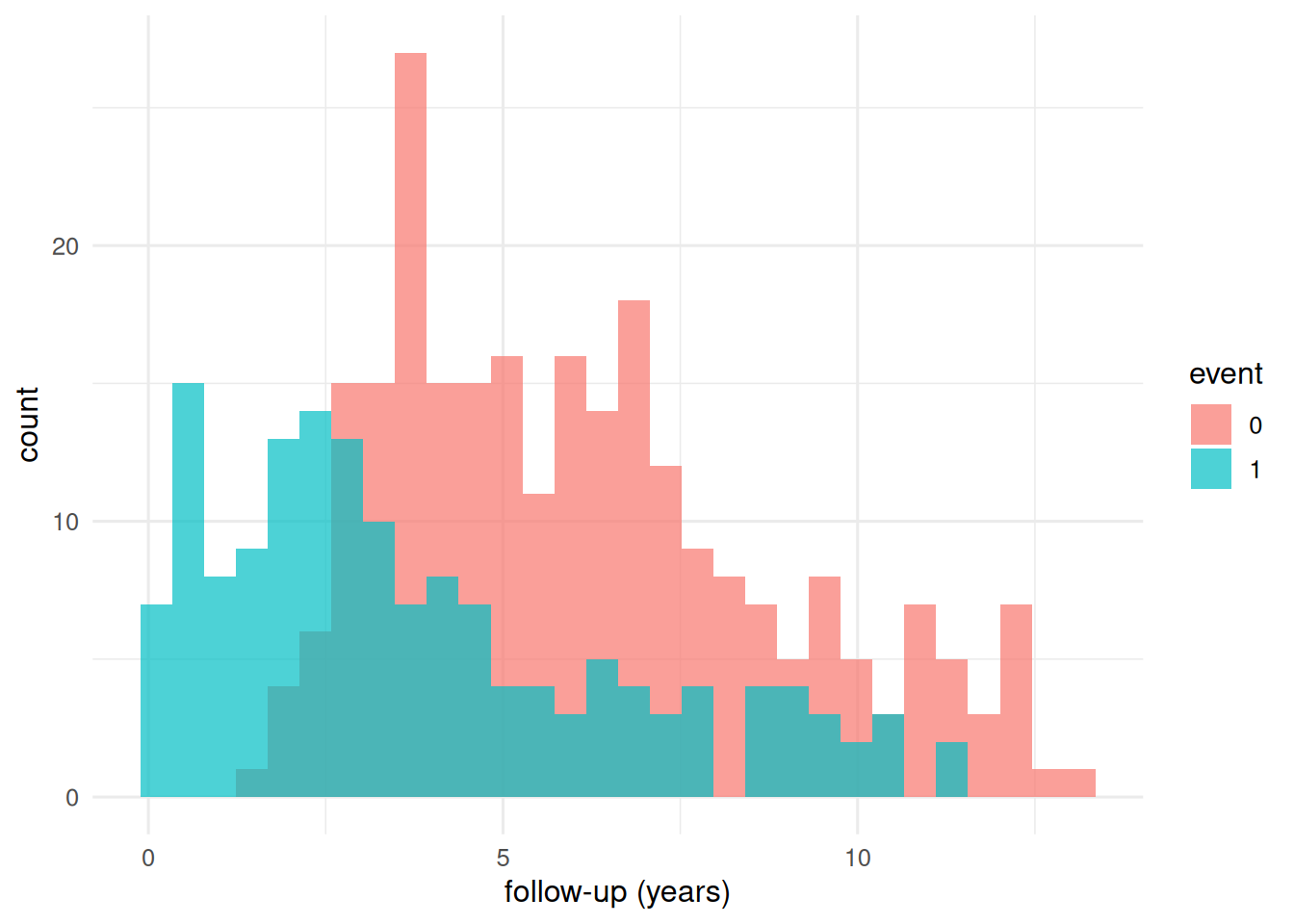
3. Execution
Sample size: 410
Number of deaths: 156
Number of trees: 500
Forest terminal node size: 15
Average no. of terminal nodes: 19.574
No. of variables tried at each split: 3
Total no. of variables: 6
Resampling used to grow trees: swor
Resample size used to grow trees: 259
Analysis: RSF
Family: surv
Splitting rule: logrank *random*
Number of random split points: 10
(OOB) CRPS: 505.07481851
(OOB) standardized CRPS: 0.12051415
(OOB) Requested performance error: 0.18039411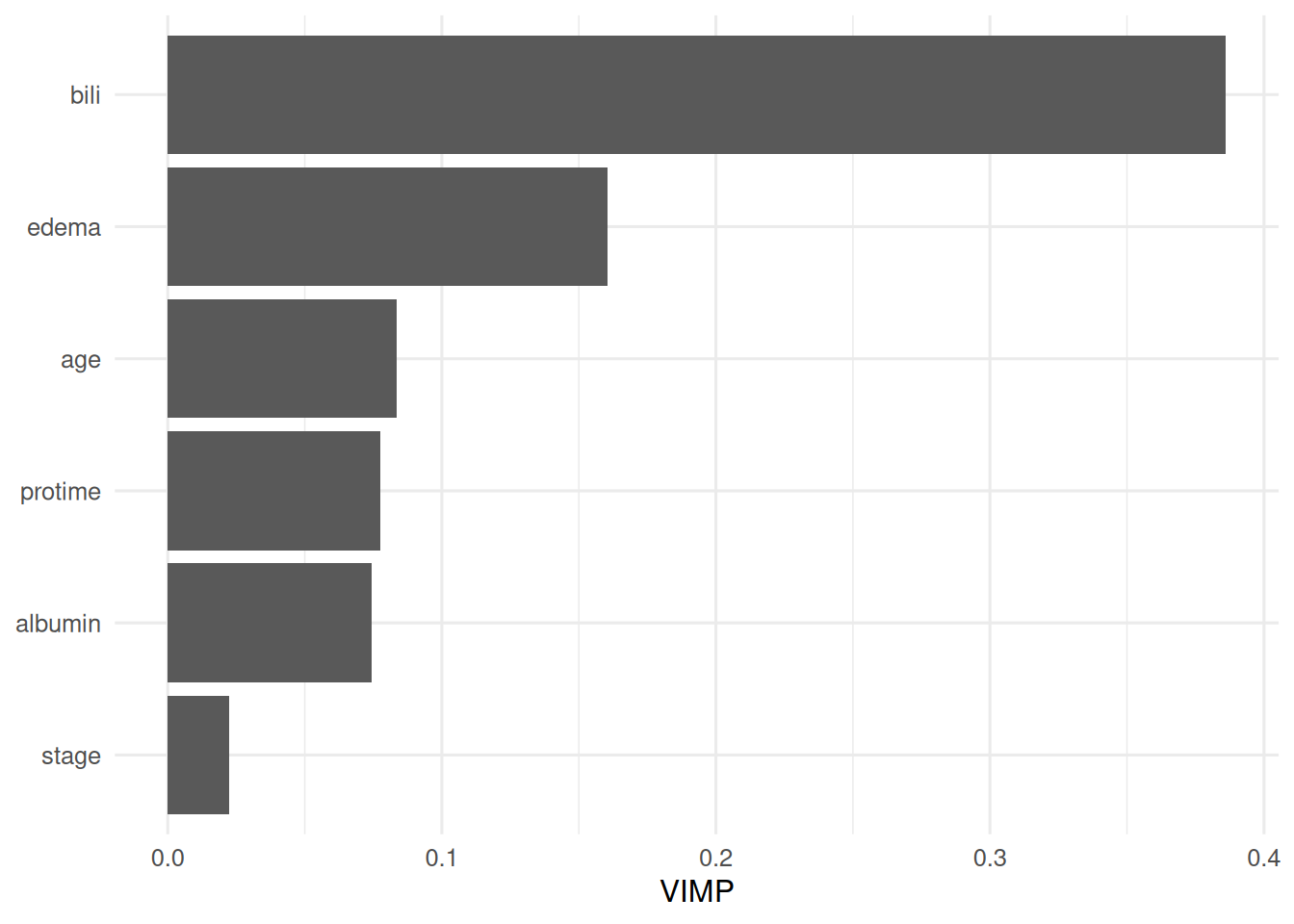
4. Check
Compare concordance on held-out data.
idx <- sample(nrow(d), 0.7 * nrow(d))
tr <- d[idx, ]; te <- d[-idx, ]
cox2 <- coxph(Surv(time, status) ~ bili + albumin + age + edema +
protime + stage, data = tr)
rsf2 <- rfsrc(Surv(time, status) ~ bili + albumin + age + edema +
protime + stage,
data = as.data.frame(tr), ntree = 500)
c_cox <- survival::concordance(cox2, newdata = te)$concordance
p_rsf <- predict(rsf2, newdata = as.data.frame(te))$predicted
c_rsf <- survival::concordance(Surv(te$time, te$status) ~ p_rsf,
reverse = TRUE)$concordance
c(cox = c_cox, rsf = c_rsf) cox rsf
0.7854077 0.8071531 DeepSurv conceptual sketch.
library(torch)
# A DeepSurv-style loss computes the partial log-likelihood on the
# sorted risk set. The network output h(x) replaces the Cox linear
# predictor; the loss is -(sum_{i:event} h(x_i) - log sum_{j in R_i} exp(h(x_j))).
deepsurv_loss <- function(risk, time, event) {
ord <- order(time, decreasing = TRUE)
risk <- risk[ord]; event <- event[ord]
log_cumsum <- torch_logcumsumexp(risk, dim = 1)
-torch_sum(event * (risk - log_cumsum)) / torch_sum(event)
}5. Report
On
survival::pbc, a random survival forest achieved test concordance 0.807 compared with 0.785 for a multivariable Cox model. Bilirubin, albumin, and age dominated VIMP. A DeepSurv network with the partial-likelihood loss would be the natural next step on a larger cohort.
On this sample size, the RSF and Cox model are close; on larger datasets with nonlinearities, the forest often pulls ahead.
Common pitfalls
- Treating the Cox proportional-hazards assumption as always met; it often is not, and the RSF is more robust to violation.
- Comparing models without the same censoring distribution in the test set.
- Reporting only C without a calibration or decision curve at the horizon of interest.
Further reading
- Ishwaran H et al. (2008), Random survival forests.
- Katzman JL et al. (2018), DeepSurv: Personalized treatment recommender system using a Cox proportional hazards deep neural network.
Session info
R version 4.4.1 (2024-06-14)
Platform: x86_64-pc-linux-gnu
Running under: Ubuntu 24.04.4 LTS
Matrix products: default
BLAS: /usr/lib/x86_64-linux-gnu/openblas-pthread/libblas.so.3
LAPACK: /usr/lib/x86_64-linux-gnu/openblas-pthread/libopenblasp-r0.3.26.so; LAPACK version 3.12.0
locale:
[1] LC_CTYPE=C.UTF-8 LC_NUMERIC=C LC_TIME=C.UTF-8
[4] LC_COLLATE=C.UTF-8 LC_MONETARY=C.UTF-8 LC_MESSAGES=C.UTF-8
[7] LC_PAPER=C.UTF-8 LC_NAME=C LC_ADDRESS=C
[10] LC_TELEPHONE=C LC_MEASUREMENT=C.UTF-8 LC_IDENTIFICATION=C
time zone: UTC
tzcode source: system (glibc)
attached base packages:
[1] stats graphics grDevices utils datasets methods base
other attached packages:
[1] randomForestSRC_3.6.2 survival_3.6-4 lubridate_1.9.5
[4] forcats_1.0.1 stringr_1.6.0 dplyr_1.2.1
[7] purrr_1.2.2 readr_2.2.0 tidyr_1.3.2
[10] tibble_3.3.1 ggplot2_4.0.3 tidyverse_2.0.0
loaded via a namespace (and not attached):
[1] Matrix_1.7-0 gtable_0.3.6 jsonlite_2.0.0 compiler_4.4.1
[5] tidyselect_1.2.1 parallel_4.4.1 DiagrammeR_1.0.12 splines_4.4.1
[9] scales_1.4.0 yaml_2.3.12 fastmap_1.2.0 lattice_0.22-6
[13] R6_2.6.1 labeling_0.4.3 generics_0.1.4 knitr_1.51
[17] visNetwork_2.1.4 htmlwidgets_1.6.4 pillar_1.11.1 RColorBrewer_1.1-3
[21] tzdb_0.5.0 rlang_1.2.0 stringi_1.8.7 xfun_0.57
[25] S7_0.2.2 otel_0.2.0 timechange_0.4.0 cli_3.6.6
[29] withr_3.0.2 magrittr_2.0.5 digest_0.6.39 grid_4.4.1
[33] hms_1.1.4 data.tree_1.2.0 lifecycle_1.0.5 vctrs_0.7.3
[37] evaluate_1.0.5 glue_1.8.1 farver_2.1.2 rmarkdown_2.31
[41] tools_4.4.1 pkgconfig_2.0.3 htmltools_0.5.9