Loads synthetic mutation data (300 mutations across 50 samples and 20 cancer-related genes) suitable for oncoplot visualization, along with clinical annotation data.
Value
A list with components:
- mutations
A data.frame with columns
sample,gene,mutation_type.- clinical
A data.frame with columns
Stage,Gender,Smoking. Rownames are sample IDs.
Examples
ex <- bb_example_mutations()
head(ex$mutations)
#> sample gene mutation_type
#> 1 TCGA-021 NRAS In_Frame_Ins
#> 2 TCGA-037 PIK3CA Missense_Mutation
#> 3 TCGA-037 BRAF Splice_Site
#> 4 TCGA-031 PIK3CA Frame_Shift_Ins
#> 5 TCGA-042 RB1 Missense_Mutation
#> 6 TCGA-008 NRAS Missense_Mutation
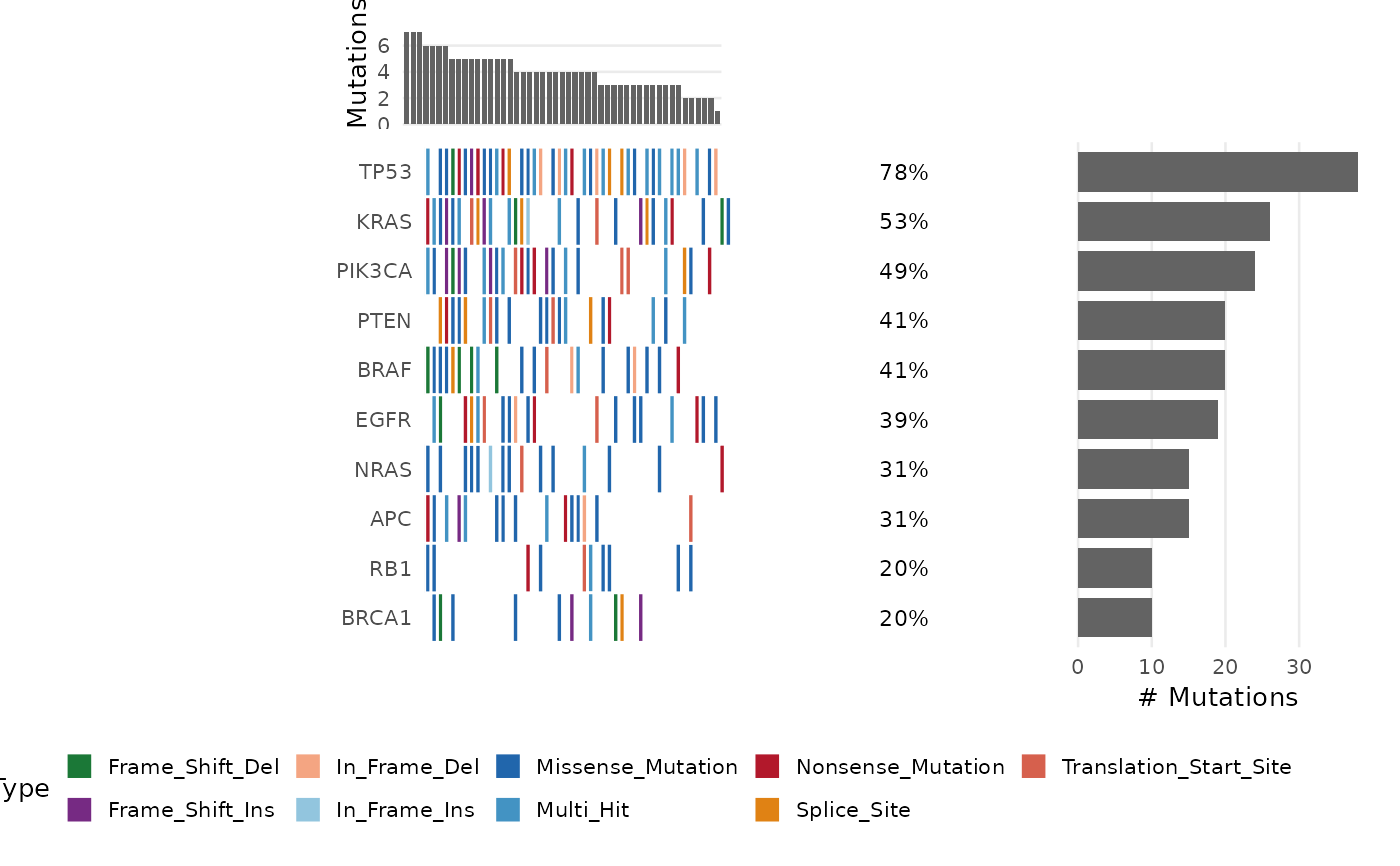
# Basic oncoplot
bb_oncoplot(ex$mutations, n_genes = 10)
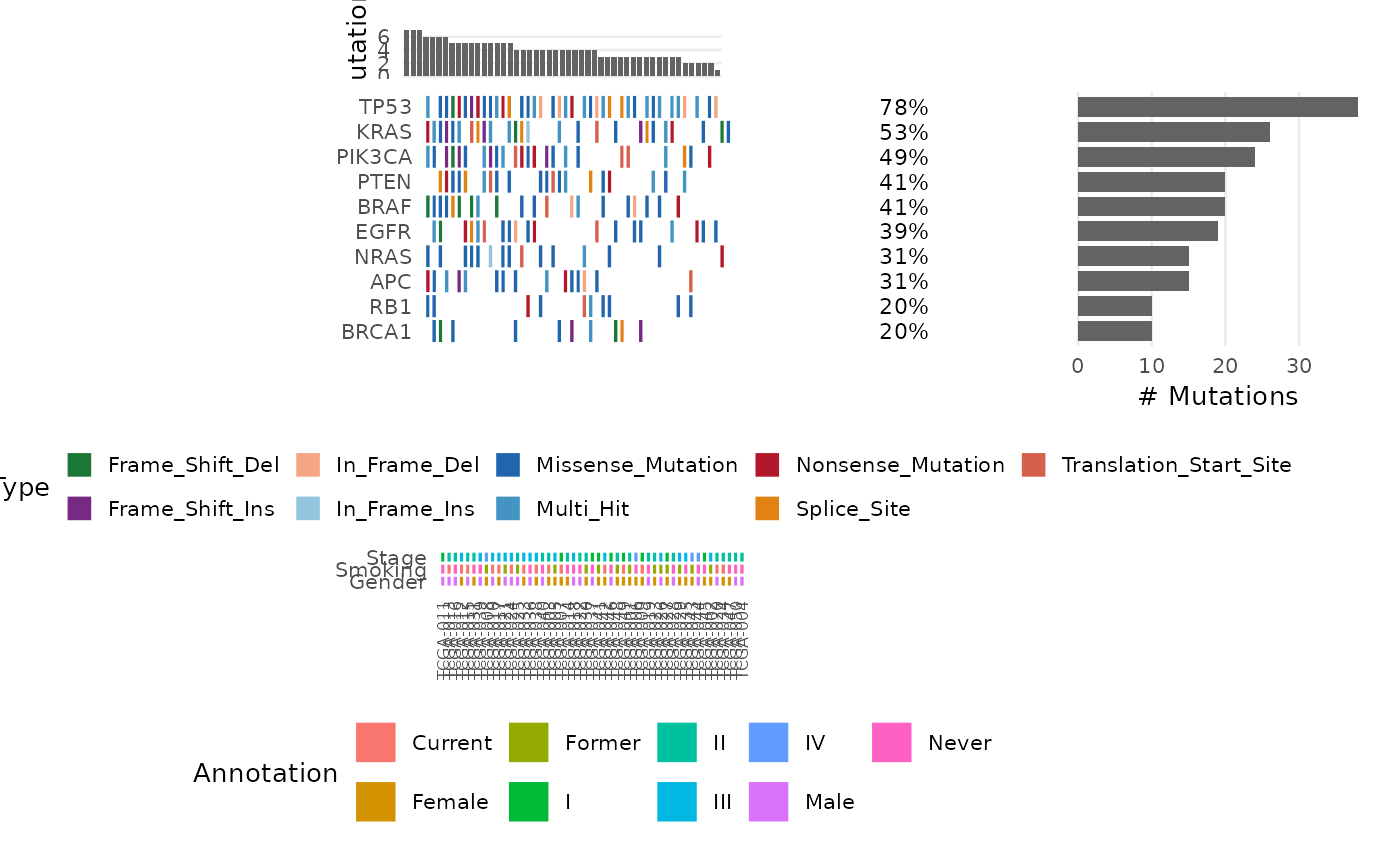
 # With clinical annotations
bb_oncoplot(ex$mutations, n_genes = 10, annotation_df = ex$clinical)
# With clinical annotations
bb_oncoplot(ex$mutations, n_genes = 10, annotation_df = ex$clinical)