library(tidyverse)
library(broom)
library(car)
library(emmeans)
set.seed(42)
theme_set(theme_minimal(base_size = 12))Week 2, Session 2 — Two-way / factorial ANOVA with interaction
Course 2 — #courses
Inference labs use the five-step template: Hypothesis → Visualise → Assumptions → Conduct → Conclude.
Learning objectives
- Fit a two-factor ANOVA and distinguish main effects from interactions.
- Visualise an interaction with
emmip()and read the picture before the table. - Report a factorial analysis in the correct order: interaction first, then conditional main effects.
Prerequisites
Session 1 of this week.
Background
A factorial design crosses two (or more) categorical factors. The two-way ANOVA decomposes variability into main effects and the interaction between factors. If the interaction is negligible, main effects can be reported as averages across levels of the other factor; if not, the main-effect table is misleading and conditional (simple) effects must be reported instead.
The canonical small dataset is ToothGrowth, which crosses dose of vitamin C with delivery method (orange juice vs ascorbic acid). It is a complete factorial and nicely balanced, which means the classification of variance into Type-I, Type-II, and Type-III sums of squares does not matter. With unbalanced designs it does, and Type-III via car::Anova with contrasts set to contr.sum is the usual safe choice.
Plotting the interaction is almost always the right first step. An interaction plot that shows parallel lines agrees with an insignificant interaction term; crossed lines signal an interaction; lines with different slopes signal a quantitative interaction.
Setup
1. Hypothesis
Does delivery method alter the effect of dose on tooth length?
Null: no interaction between supp and dose. Alternative: the dose effect differs by supp.
2. Visualise
tg <- ToothGrowth |> mutate(dose = factor(dose))
ggplot(tg, aes(dose, len, colour = supp, group = supp)) +
stat_summary(fun = mean, geom = "point", size = 3) +
stat_summary(fun = mean, geom = "line") +
stat_summary(fun.data = mean_cl_normal, geom = "errorbar", width = 0.1) +
labs(x = "Dose (mg/day)", y = "Tooth length", colour = "Supp")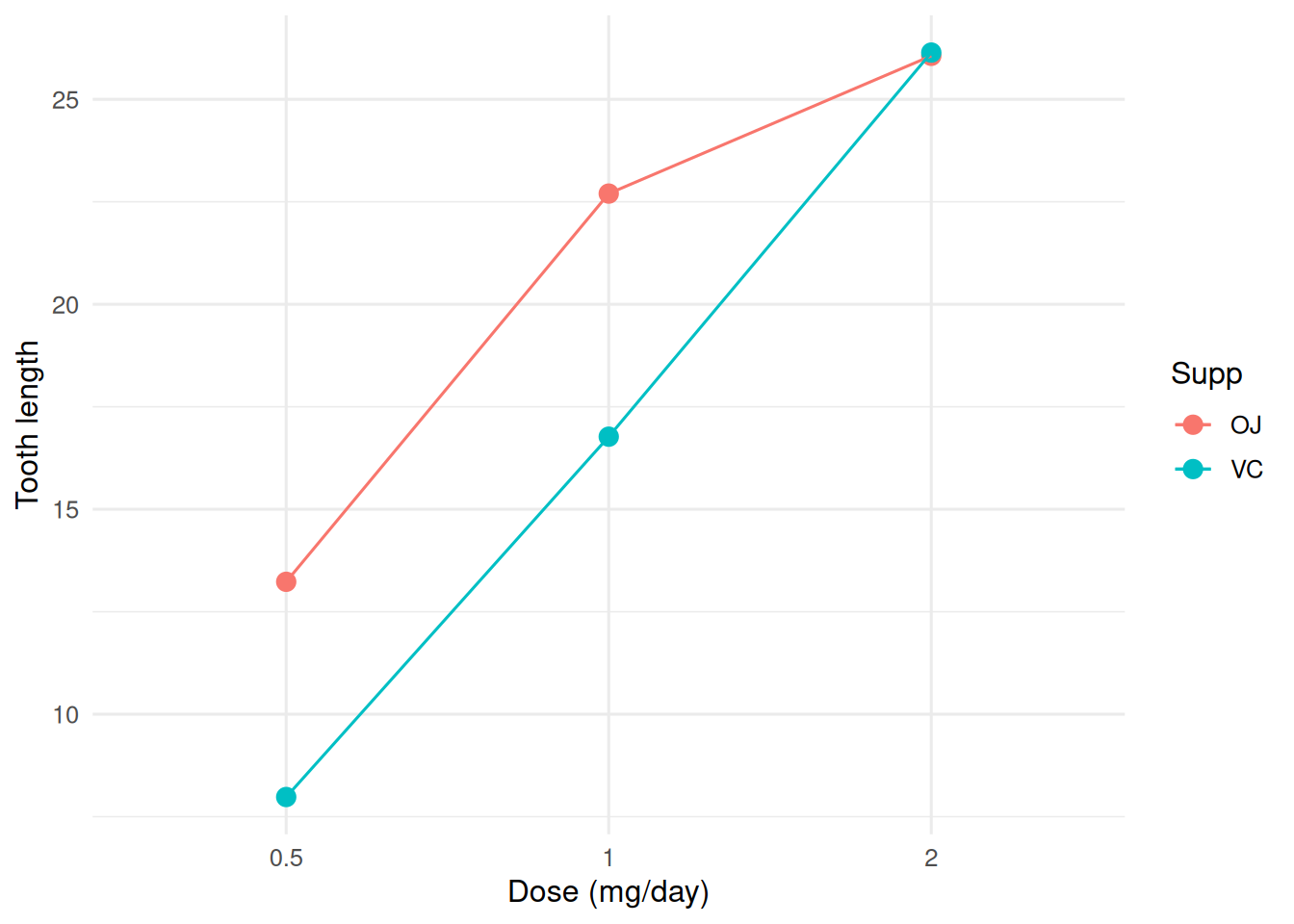
3. Assumptions
fit <- aov(len ~ supp * dose, data = tg)
par(mfrow = c(2, 2))
plot(fit)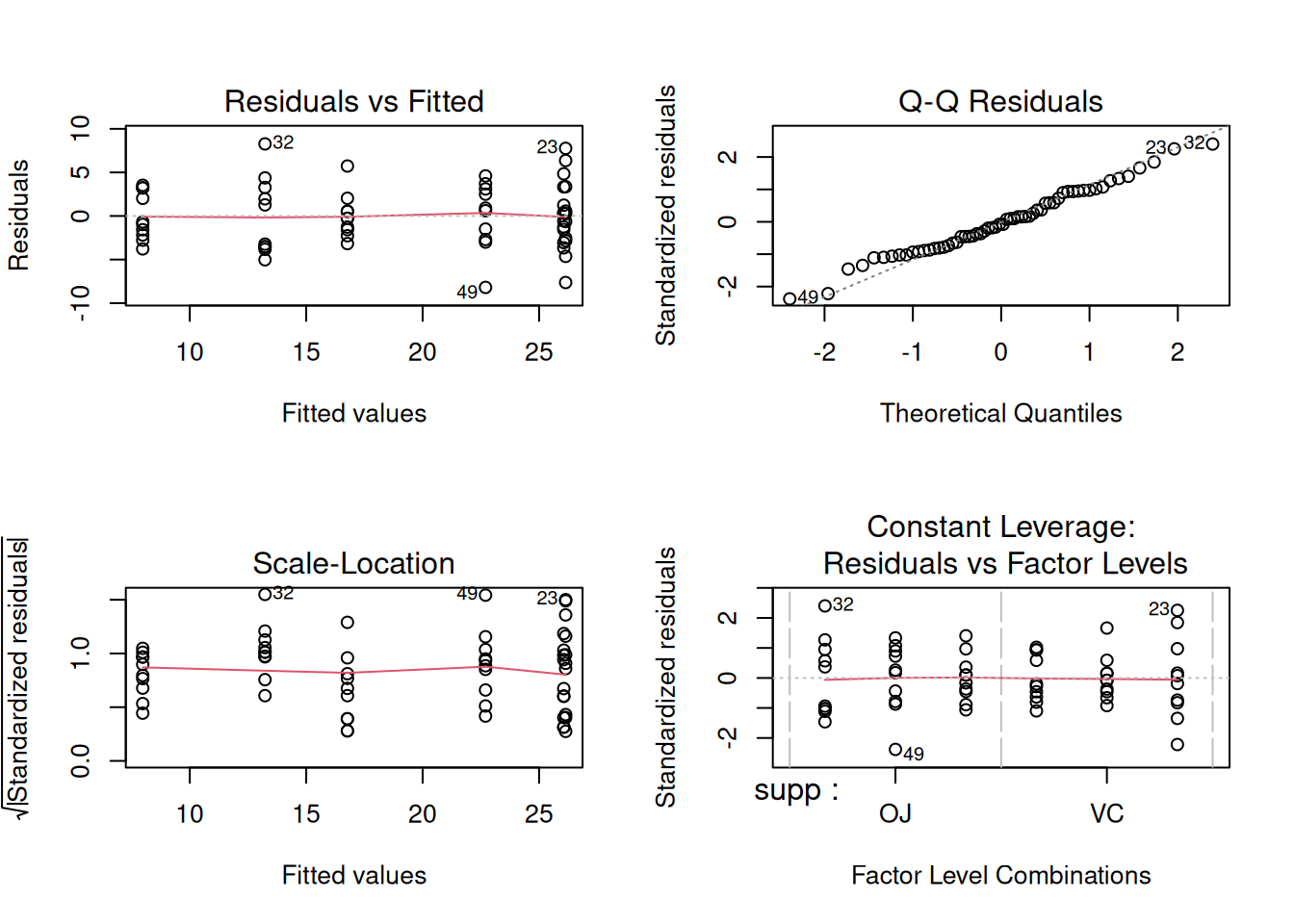
par(mfrow = c(1, 1))4. Conduct
# Type-II sums of squares for a balanced design
Anova(lm(len ~ supp * dose, data = tg), type = 2)Anova Table (Type II tests)
Response: len
Sum Sq Df F value Pr(>F)
supp 205.35 1 15.572 0.0002312 ***
dose 2426.43 2 92.000 < 2.2e-16 ***
supp:dose 108.32 2 4.107 0.0218603 *
Residuals 712.11 54
---
Signif. codes: 0 '***' 0.001 '**' 0.01 '*' 0.05 '.' 0.1 ' ' 1emm <- emmeans(fit, ~ supp | dose)
pairs(emm)dose = 0.5:
contrast estimate SE df t.ratio p.value
OJ - VC 5.25 1.62 54 3.233 0.0021
dose = 1:
contrast estimate SE df t.ratio p.value
OJ - VC 5.93 1.62 54 3.651 0.0006
dose = 2:
contrast estimate SE df t.ratio p.value
OJ - VC -0.08 1.62 54 -0.049 0.9609emmip(fit, supp ~ dose, CIs = TRUE)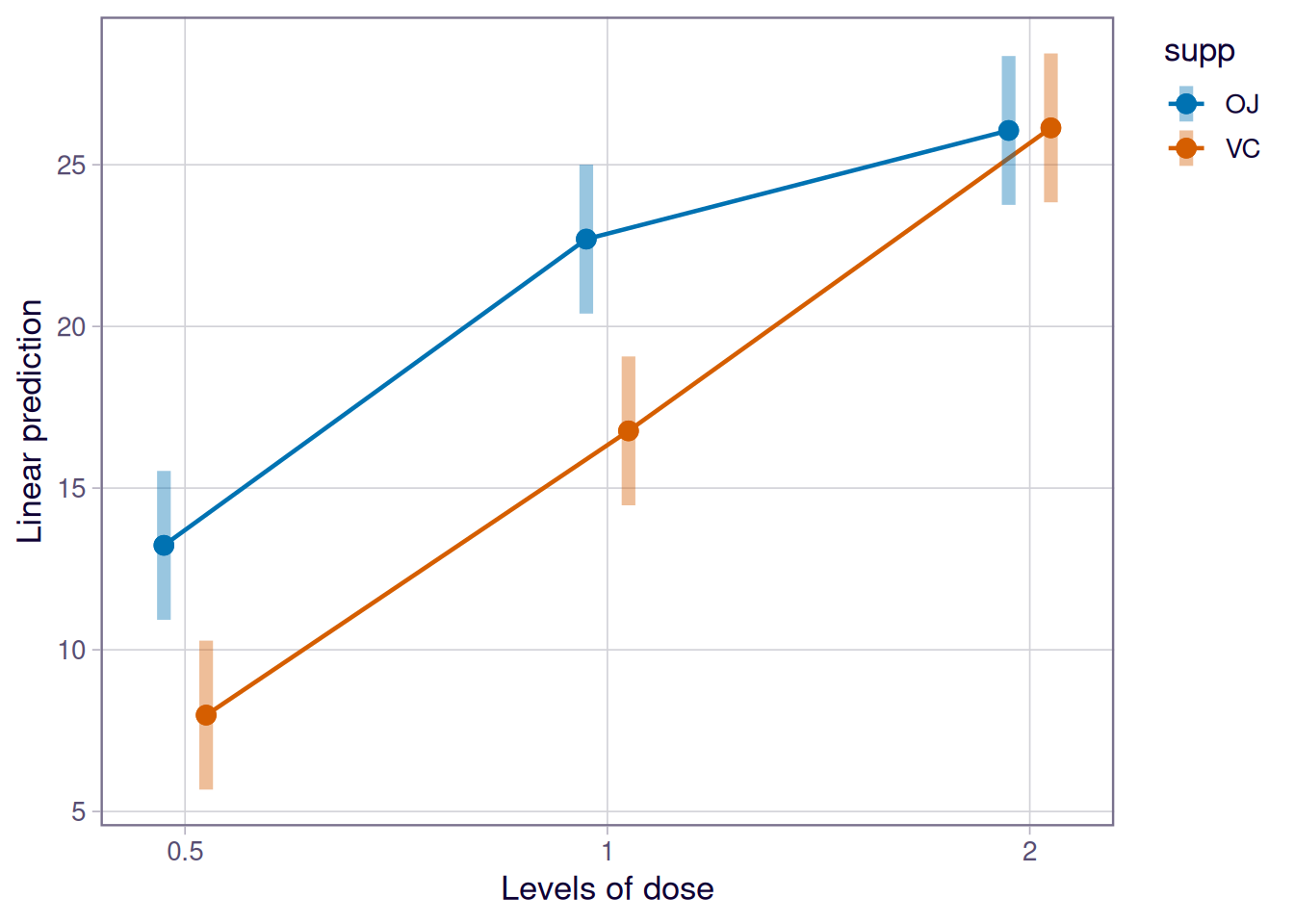
5. Concluding statement
Tooth length responded to dose (large main effect) and to supplement (smaller main effect), with evidence of an interaction (see ANOVA table). The difference between orange juice and ascorbic acid was largest at the lowest dose and negligible at the highest.
Use the interaction plot to justify the order of the report: when the lines are not parallel, the main-effect column in the ANOVA table is only a summary, not the headline.
Common pitfalls
- Reporting main effects without first checking the interaction.
- Using Type-III sums of squares with default treatment contrasts in unbalanced designs; the output depends on the contrast coding.
- Ignoring unequal cell sizes when reading an ANOVA table.
Further reading
- Fox J, Weisberg S. An R Companion to Applied Regression, ch. 5.
- Cochran WG, Cox GM. Experimental Designs.
- Wilkinson L, Rogers CE (1973), Symbolic description of factorial…
Session info
sessionInfo()R version 4.4.1 (2024-06-14)
Platform: x86_64-pc-linux-gnu
Running under: Ubuntu 24.04.4 LTS
Matrix products: default
BLAS: /usr/lib/x86_64-linux-gnu/openblas-pthread/libblas.so.3
LAPACK: /usr/lib/x86_64-linux-gnu/openblas-pthread/libopenblasp-r0.3.26.so; LAPACK version 3.12.0
locale:
[1] LC_CTYPE=C.UTF-8 LC_NUMERIC=C LC_TIME=C.UTF-8
[4] LC_COLLATE=C.UTF-8 LC_MONETARY=C.UTF-8 LC_MESSAGES=C.UTF-8
[7] LC_PAPER=C.UTF-8 LC_NAME=C LC_ADDRESS=C
[10] LC_TELEPHONE=C LC_MEASUREMENT=C.UTF-8 LC_IDENTIFICATION=C
time zone: UTC
tzcode source: system (glibc)
attached base packages:
[1] stats graphics grDevices utils datasets methods base
other attached packages:
[1] emmeans_2.0.3 car_3.1-5 carData_3.0-6 broom_1.0.12
[5] lubridate_1.9.5 forcats_1.0.1 stringr_1.6.0 dplyr_1.2.1
[9] purrr_1.2.2 readr_2.2.0 tidyr_1.3.2 tibble_3.3.1
[13] ggplot2_4.0.3 tidyverse_2.0.0
loaded via a namespace (and not attached):
[1] generics_0.1.4 stringi_1.8.7 lattice_0.22-6 hms_1.1.4
[5] digest_0.6.39 magrittr_2.0.5 evaluate_1.0.5 grid_4.4.1
[9] timechange_0.4.0 estimability_1.5.1 RColorBrewer_1.1-3 mvtnorm_1.3-7
[13] fastmap_1.2.0 jsonlite_2.0.0 backports_1.5.1 Formula_1.2-5
[17] scales_1.4.0 abind_1.4-8 cli_3.6.6 rlang_1.2.0
[21] withr_3.0.2 yaml_2.3.12 otel_0.2.0 tools_4.4.1
[25] tzdb_0.5.0 coda_0.19-4.1 vctrs_0.7.3 R6_2.6.1
[29] lifecycle_1.0.5 htmlwidgets_1.6.4 pkgconfig_2.0.3 pillar_1.11.1
[33] gtable_0.3.6 glue_1.8.1 xfun_0.57 tidyselect_1.2.1
[37] knitr_1.51 xtable_1.8-8 farver_2.1.2 htmltools_0.5.9
[41] labeling_0.4.3 rmarkdown_2.31 compiler_4.4.1 S7_0.2.2