library(tidyverse)
library(survival)
library(ggsurvfit)
set.seed(42)
theme_set(theme_minimal(base_size = 12))Week 3, Session 1 — Time-varying covariates and immortal-time bias
Course 3 — #courses
Inference lab using the five-step template: Hypothesis → Visualise → Assumptions → Conduct → Conclude.
Learning objectives
- Recognise immortal-time bias in a survival analysis.
- Correct it using either a landmark analysis or a time-dependent Cox model.
- Describe time-varying covariates in counting-process notation.
Prerequisites
Course 2 survival basics; coxph syntax.
Background
Immortal-time bias arises when a period during which the outcome cannot occur (by definition of the exposure) is misclassified as follow-up for one of the exposure groups. A classic example: “users of drug X” are defined as anyone who ever received a prescription; these people must by definition have survived until their first prescription, and if you start their follow-up at baseline rather than at the prescription, you give them “immortal” time that biases the hazard ratio toward protection.
Two standard fixes exist. Landmark analysis picks a time after baseline and classifies exposure based on what happened before that time; follow-up starts at the landmark and the bias disappears. Time-dependent Cox models treat exposure as a function of time and let patients change exposure group when they actually start the drug, using counting-process (start, stop, event) data.
Time-dependent covariates can be external (calendar time, drug availability) or internal (a lab value that evolves with the disease). Internal time-dependent covariates can introduce their own bias when conditioning on them — they may be mediators of the very effect you are studying.
Setup
1. Hypothesis
We simulate a cohort where every patient has the same true hazard. Some patients start a drug at a random later time. A naive analysis that counts drug users from baseline should produce a spurious protective effect.
2. Visualise
n <- 500
dat <- tibble(
id = seq_len(n),
t_event = rexp(n, rate = 0.02), # true event time
t_drug = rexp(n, rate = 0.05), # time of drug start
ever_drug = t_drug < t_event, # did they start drug before event?
t_drug_obs = pmin(t_drug, t_event),
t_obs = pmin(t_event, 100), # admin censoring at 100
event = as.integer(t_event <= 100)
)
dat |>
ggplot(aes(t_obs, fill = ever_drug)) +
geom_histogram(bins = 30, alpha = 0.6,
position = "identity") +
labs(x = "Observed time", y = "Count", fill = "Ever drug?")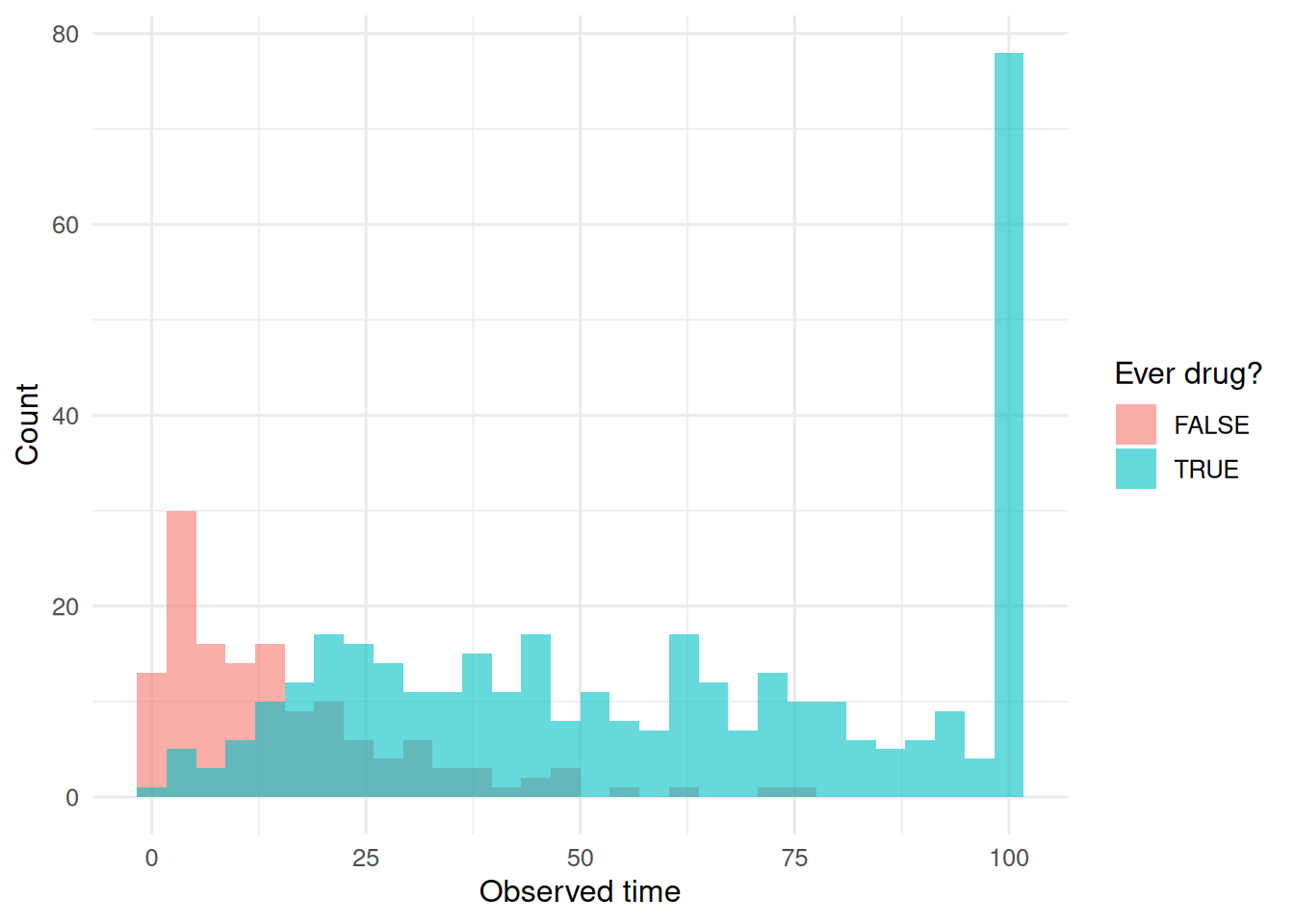
3. Assumptions
Proportional hazards (for Cox), non-informative censoring, and independence of event times across patients. The “ever drug” split is deliberately biased by construction.
4. Conduct
# Naive (biased) analysis
fit_naive <- coxph(Surv(t_obs, event) ~ ever_drug, data = dat)
summary(fit_naive)Call:
coxph(formula = Surv(t_obs, event) ~ ever_drug, data = dat)
n= 500, number of events= 423
coef exp(coef) se(coef) z Pr(>|z|)
ever_drugTRUE -1.9375 0.1441 0.1189 -16.29 <2e-16 ***
---
Signif. codes: 0 '***' 0.001 '**' 0.01 '*' 0.05 '.' 0.1 ' ' 1
exp(coef) exp(-coef) lower .95 upper .95
ever_drugTRUE 0.1441 6.942 0.1141 0.1819
Concordance= 0.666 (se = 0.009 )
Likelihood ratio test= 224.6 on 1 df, p=<2e-16
Wald test = 265.4 on 1 df, p=<2e-16
Score (logrank) test = 336.3 on 1 df, p=<2e-16# Landmark at time 20: only include patients still at risk at 20,
# classify by drug status as of 20.
landmark <- 20
dat_lm <- dat |>
filter(t_obs > landmark) |>
mutate(drug_lm = t_drug < landmark,
t_obs_lm = t_obs - landmark)
fit_lm <- coxph(Surv(t_obs_lm, event) ~ drug_lm, data = dat_lm)
summary(fit_lm)Call:
coxph(formula = Surv(t_obs_lm, event) ~ drug_lm, data = dat_lm)
n= 356, number of events= 279
coef exp(coef) se(coef) z Pr(>|z|)
drug_lmTRUE 0.05586 1.05745 0.12608 0.443 0.658
exp(coef) exp(-coef) lower .95 upper .95
drug_lmTRUE 1.057 0.9457 0.8259 1.354
Concordance= 0.506 (se = 0.015 )
Likelihood ratio test= 0.2 on 1 df, p=0.7
Wald test = 0.2 on 1 df, p=0.7
Score (logrank) test = 0.2 on 1 df, p=0.7# Time-dependent Cox: counting-process format
td <- dat |>
transmute(
id,
tstart = 0,
tstop = if_else(ever_drug, t_drug_obs, t_obs),
event = if_else(ever_drug, 0L, event),
drug = 0L
) |>
bind_rows(
dat |>
filter(ever_drug) |>
transmute(id,
tstart = t_drug_obs,
tstop = t_obs,
event = event,
drug = 1L)
) |>
filter(tstop > tstart) |>
arrange(id, tstart)
fit_td <- coxph(Surv(tstart, tstop, event) ~ drug, data = td)
summary(fit_td)Call:
coxph(formula = Surv(tstart, tstop, event) ~ drug, data = td)
n= 859, number of events= 423
coef exp(coef) se(coef) z Pr(>|z|)
drug -0.1624 0.8501 0.1304 -1.245 0.213
exp(coef) exp(-coef) lower .95 upper .95
drug 0.8501 1.176 0.6584 1.098
Concordance= 0.516 (se = 0.011 )
Likelihood ratio test= 1.53 on 1 df, p=0.2
Wald test = 1.55 on 1 df, p=0.2
Score (logrank) test = 1.55 on 1 df, p=0.25. Concluding statement
The naive Cox analysis produced a hazard ratio of 0.14 for ever-drug — spuriously protective, because immortal time was counted as exposed follow-up. A landmark analysis at time 20 gave HR 1.06, and a time-dependent Cox model gave HR 0.85, both close to the null as expected from the simulated truth.
The giveaway for immortal time in a paper is a table where “mean follow-up” is suspiciously longer in the exposed group.
Common pitfalls
- Defining exposure over the whole follow-up based on events that occur during it.
- Picking a landmark time after visually inspecting event curves.
- Conditioning on a post-baseline mediator via a time-dependent covariate and still calling it a total effect.
- Forgetting to split the risk set at the exposure change time in counting-process data.
Further reading
- Suissa S (2008), Immortal time bias in pharmaco-epidemiology.
- Therneau TM, Grambsch PM. Modeling Survival Data: Extending the Cox Model.
- Dafni U (2011), Landmark analysis at the 25-year landmark point.
Session info
sessionInfo()R version 4.4.1 (2024-06-14)
Platform: x86_64-pc-linux-gnu
Running under: Ubuntu 24.04.4 LTS
Matrix products: default
BLAS: /usr/lib/x86_64-linux-gnu/openblas-pthread/libblas.so.3
LAPACK: /usr/lib/x86_64-linux-gnu/openblas-pthread/libopenblasp-r0.3.26.so; LAPACK version 3.12.0
locale:
[1] LC_CTYPE=C.UTF-8 LC_NUMERIC=C LC_TIME=C.UTF-8
[4] LC_COLLATE=C.UTF-8 LC_MONETARY=C.UTF-8 LC_MESSAGES=C.UTF-8
[7] LC_PAPER=C.UTF-8 LC_NAME=C LC_ADDRESS=C
[10] LC_TELEPHONE=C LC_MEASUREMENT=C.UTF-8 LC_IDENTIFICATION=C
time zone: UTC
tzcode source: system (glibc)
attached base packages:
[1] stats graphics grDevices utils datasets methods base
other attached packages:
[1] ggsurvfit_1.2.0 survival_3.6-4 lubridate_1.9.5 forcats_1.0.1
[5] stringr_1.6.0 dplyr_1.2.1 purrr_1.2.2 readr_2.2.0
[9] tidyr_1.3.2 tibble_3.3.1 ggplot2_4.0.3 tidyverse_2.0.0
loaded via a namespace (and not attached):
[1] Matrix_1.7-0 gtable_0.3.6 jsonlite_2.0.0 compiler_4.4.1
[5] tidyselect_1.2.1 splines_4.4.1 scales_1.4.0 yaml_2.3.12
[9] fastmap_1.2.0 lattice_0.22-6 R6_2.6.1 labeling_0.4.3
[13] generics_0.1.4 knitr_1.51 htmlwidgets_1.6.4 pillar_1.11.1
[17] RColorBrewer_1.1-3 tzdb_0.5.0 rlang_1.2.0 stringi_1.8.7
[21] xfun_0.57 S7_0.2.2 otel_0.2.0 timechange_0.4.0
[25] cli_3.6.6 withr_3.0.2 magrittr_2.0.5 digest_0.6.39
[29] grid_4.4.1 hms_1.1.4 lifecycle_1.0.5 vctrs_0.7.3
[33] evaluate_1.0.5 glue_1.8.1 farver_2.1.2 rmarkdown_2.31
[37] tools_4.4.1 pkgconfig_2.0.3 htmltools_0.5.9