library(tidyverse)
library(survival)
library(ggsurvfit)
library(broom)
set.seed(42)
theme_set(theme_minimal(base_size = 12))Week 4, Session 3 — Survival primer: KM, log-rank, Cox PH
Course 2 — #courses
Inference lab: Hypothesis → Visualise → Assumptions → Conduct → Conclude.
Learning objectives
- Estimate and plot a Kaplan-Meier survival curve for a two-group comparison.
- Run and interpret a log-rank test.
- Fit a Cox proportional-hazards model, check the PH assumption, and read a hazard ratio honestly.
Prerequisites
Comfort with GLMs and with reading a regression table.
Background
Survival analysis handles outcomes that are times to an event, where some subjects have not yet had the event when observation ends — the familiar right-censoring problem. Ignoring censoring by throwing out the censored observations biases the estimator; pretending the censored observations are event-free biases it the other way. The survival machinery handles censoring correctly by reasoning at each event time about who is still at risk.
Three tools cover most clinical applications: the Kaplan-Meier estimator for a non-parametric survival curve, the log-rank test for a two-group comparison, and Cox proportional-hazards regression for adjustment and multivariable modelling. The PH assumption — that the hazard ratio is constant over time — deserves explicit checking; when it fails, a time-varying coefficient or a stratified model is usually the right fix.
Setup
1. Hypothesis
Using survival::lung, compare survival between male and female patients. H₀: hazard ratio = 1. H₁: hazard ratio ≠ 1. α = 0.05.
2. Visualise
lung2 <- lung |>
mutate(sex = factor(sex, levels = 1:2, labels = c("male", "female")))
km <- survfit2(Surv(time, status) ~ sex, data = lung2)
km |>
ggsurvfit() +
add_confidence_interval() +
add_risktable() +
labs(x = "Days", y = "Survival probability")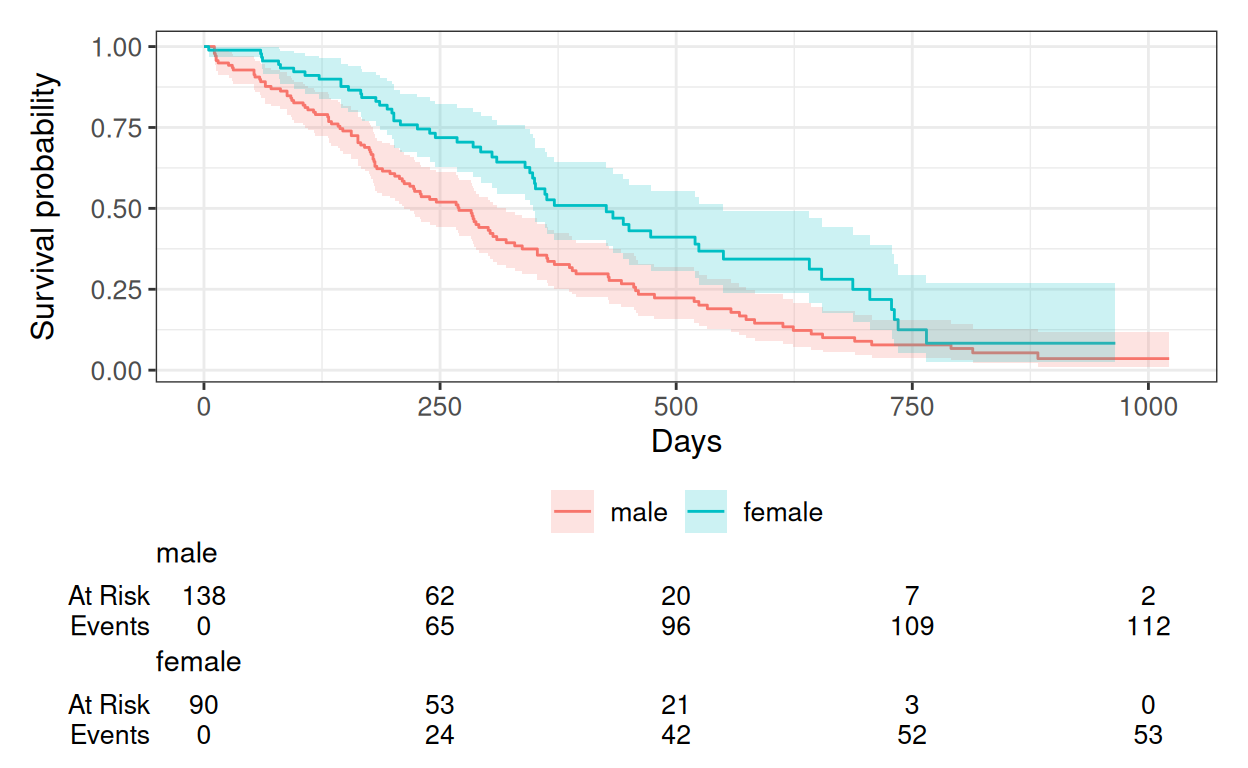
3. Assumptions
Two assumptions are worth checking: non-informative censoring (patients censored at a given time are representative of those still at risk) and proportional hazards.
cox_fit <- coxph(Surv(time, status) ~ sex + age, data = lung2)
cox.zph(cox_fit) chisq df p
sex 2.608 1 0.11
age 0.209 1 0.65
GLOBAL 2.771 2 0.25A large p-value on each row and a non-systematic Schoenfeld residual plot support the PH assumption; a small p calls for a time-varying coefficient or stratification.
4. Conduct
lr <- survdiff(Surv(time, status) ~ sex, data = lung2)
lrCall:
survdiff(formula = Surv(time, status) ~ sex, data = lung2)
N Observed Expected (O-E)^2/E (O-E)^2/V
sex=male 138 112 91.6 4.55 10.3
sex=female 90 53 73.4 5.68 10.3
Chisq= 10.3 on 1 degrees of freedom, p= 0.001 tidy(cox_fit, exponentiate = TRUE, conf.int = TRUE)# A tibble: 2 × 7
term estimate std.error statistic p.value conf.low conf.high
<chr> <dbl> <dbl> <dbl> <dbl> <dbl> <dbl>
1 sexfemale 0.599 0.167 -3.06 0.00218 0.431 0.831
2 age 1.02 0.00922 1.85 0.0646 0.999 1.04 5. Concluding statement
Female sex was associated with a lower risk of death relative to male sex (HR 0.6; 95% CI 0.43 to 0.83; p = 0.0022), adjusted for age. The proportional-hazards assumption was not clearly violated (Schoenfeld test, all p > 0.05).
Emphasise the difference between a median survival comparison (which requires enough events) and a Cox model (which uses every event time).
Common pitfalls
- Reporting a median survival that is not reached and calling it “undefined” in the results. Say so explicitly.
- Ignoring the PH check because the output looks clean.
- Comparing Kaplan-Meier curves by eye alone — add the log-rank test and the risk table.
- Collapsing a competing event (e.g. death from another cause) into censoring without justification.
Further reading
- Harrell FE. Regression Modeling Strategies, ch. 20.
- Therneau TM & Grambsch PM. Modeling Survival Data.
Session info
sessionInfo()R version 4.4.1 (2024-06-14)
Platform: x86_64-pc-linux-gnu
Running under: Ubuntu 24.04.4 LTS
Matrix products: default
BLAS: /usr/lib/x86_64-linux-gnu/openblas-pthread/libblas.so.3
LAPACK: /usr/lib/x86_64-linux-gnu/openblas-pthread/libopenblasp-r0.3.26.so; LAPACK version 3.12.0
locale:
[1] LC_CTYPE=C.UTF-8 LC_NUMERIC=C LC_TIME=C.UTF-8
[4] LC_COLLATE=C.UTF-8 LC_MONETARY=C.UTF-8 LC_MESSAGES=C.UTF-8
[7] LC_PAPER=C.UTF-8 LC_NAME=C LC_ADDRESS=C
[10] LC_TELEPHONE=C LC_MEASUREMENT=C.UTF-8 LC_IDENTIFICATION=C
time zone: UTC
tzcode source: system (glibc)
attached base packages:
[1] stats graphics grDevices utils datasets methods base
other attached packages:
[1] broom_1.0.12 ggsurvfit_1.2.0 survival_3.6-4 lubridate_1.9.5
[5] forcats_1.0.1 stringr_1.6.0 dplyr_1.2.1 purrr_1.2.2
[9] readr_2.2.0 tidyr_1.3.2 tibble_3.3.1 ggplot2_4.0.3
[13] tidyverse_2.0.0
loaded via a namespace (and not attached):
[1] Matrix_1.7-0 gtable_0.3.6 jsonlite_2.0.0 compiler_4.4.1
[5] tidyselect_1.2.1 splines_4.4.1 scales_1.4.0 yaml_2.3.12
[9] fastmap_1.2.0 lattice_0.22-6 R6_2.6.1 patchwork_1.3.2
[13] labeling_0.4.3 generics_0.1.4 knitr_1.51 backports_1.5.1
[17] htmlwidgets_1.6.4 pillar_1.11.1 RColorBrewer_1.1-3 tzdb_0.5.0
[21] rlang_1.2.0 utf8_1.2.6 stringi_1.8.7 xfun_0.57
[25] S7_0.2.2 otel_0.2.0 timechange_0.4.0 cli_3.6.6
[29] withr_3.0.2 magrittr_2.0.5 digest_0.6.39 grid_4.4.1
[33] hms_1.1.4 lifecycle_1.0.5 vctrs_0.7.3 evaluate_1.0.5
[37] glue_1.8.1 farver_2.1.2 rmarkdown_2.31 tools_4.4.1
[41] pkgconfig_2.0.3 htmltools_0.5.9