library(tidyverse)
library(ranger)
library(DALEX)
library(iml)
set.seed(42)
theme_set(theme_minimal(base_size = 12))Week 2, Session 2 — Interpretability and SHAP
Course 4 — #courses
Workflow labs use the variant template: Goal → Approach → Execution → Check → Report.
Learning objectives
- Produce permutation importance, partial-dependence, and Shapley values for a fitted black-box model, and interpret each plot.
- State the limitations of feature-attribution methods under feature correlation.
- Wrap a model with
DALEXandimlfor a standardised explanation pipeline.
Prerequisites
Trees and forests from Session 1.
Background
Interpretability is not a single method; it is a cluster of tools that answer different questions. Permutation importance answers: how much worse does the model score if I scramble this feature? Partial dependence answers: averaging over all the other features, how does the prediction change as this feature varies? SHAP answers: for this specific prediction, how much did each feature contribute relative to the average prediction?
All three can mislead under feature correlation. Permuting a feature breaks its correlation with others, producing unrealistic inputs. Partial dependence averages over a marginal distribution that may not represent the data. SHAP — in particular TreeSHAP for trees — has several variants (interventional vs conditional) that answer subtly different causal questions. The robust practice is to use more than one method and to check that they tell a consistent story.
Interpretability is not a substitute for a causal model. A feature can have large SHAP values and large permutation importance and still be downstream of the outcome, a proxy for a confounder, or a leakage artefact. The explanation tells you what the model uses; whether that is what the world uses is a separate, harder question.
Setup
1. Goal
Fit a random forest on the palmerpenguins dataset and produce three complementary interpretability artefacts.
2. Approach
d <- palmerpenguins::penguins |>
tidyr::drop_na() |>
mutate(species = factor(species))
ggplot(d, aes(bill_length_mm, body_mass_g, colour = species)) +
geom_point(alpha = 0.7)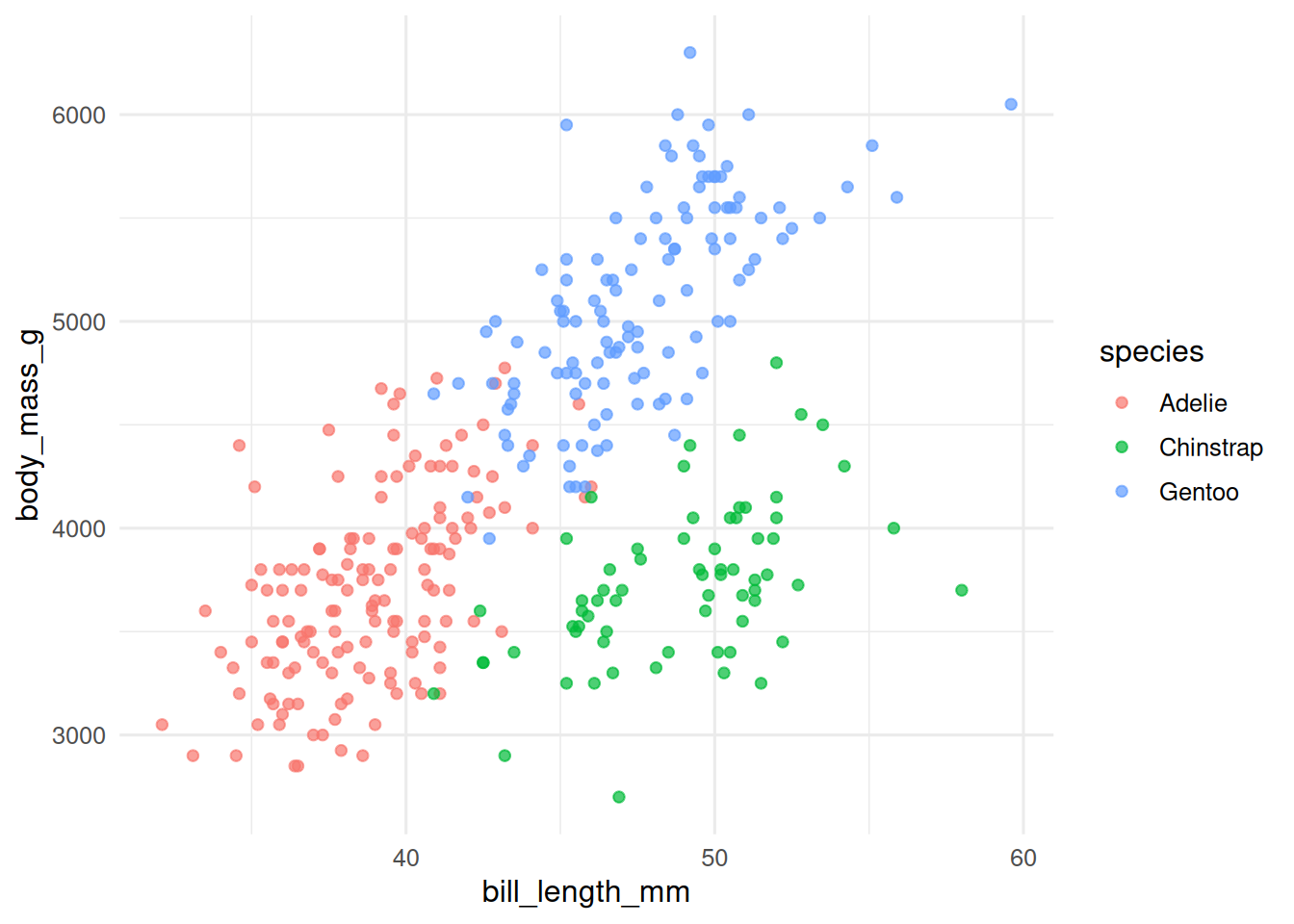
3. Execution
rf <- ranger(species ~ bill_length_mm + bill_depth_mm +
flipper_length_mm + body_mass_g,
data = d, probability = TRUE, num.trees = 500)
explainer <- DALEX::explain(
model = rf,
data = d[, c("bill_length_mm", "bill_depth_mm",
"flipper_length_mm", "body_mass_g")],
y = as.integer(d$species == "Adelie"),
predict_function = function(m, newdata) predict(m, newdata)$predictions[, "Adelie"],
label = "rf-Adelie",
verbose = FALSE
)
vi <- DALEX::model_parts(explainer, type = "variable_importance")
plot(vi)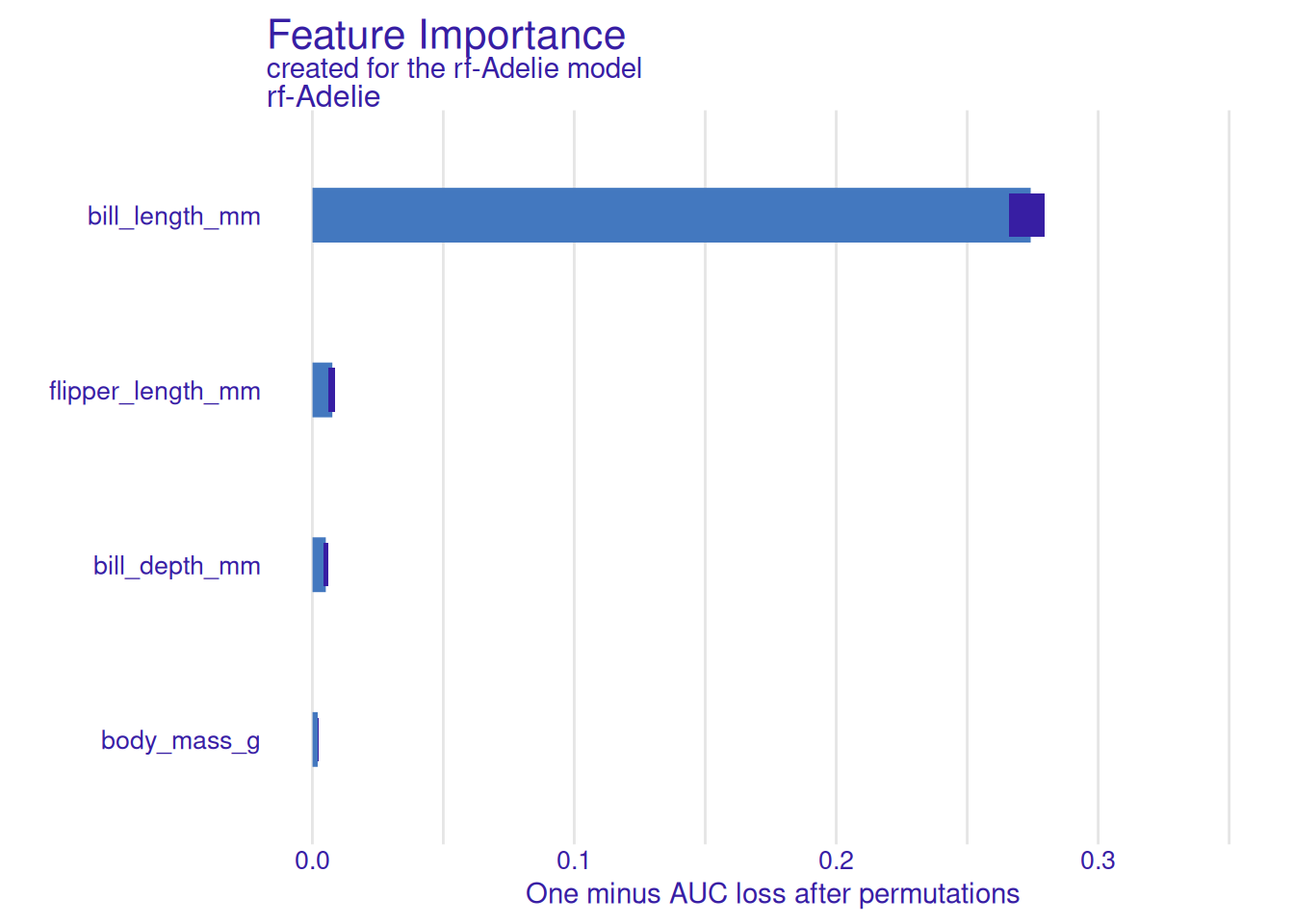
pdp <- DALEX::model_profile(explainer, variables = "bill_length_mm")
plot(pdp)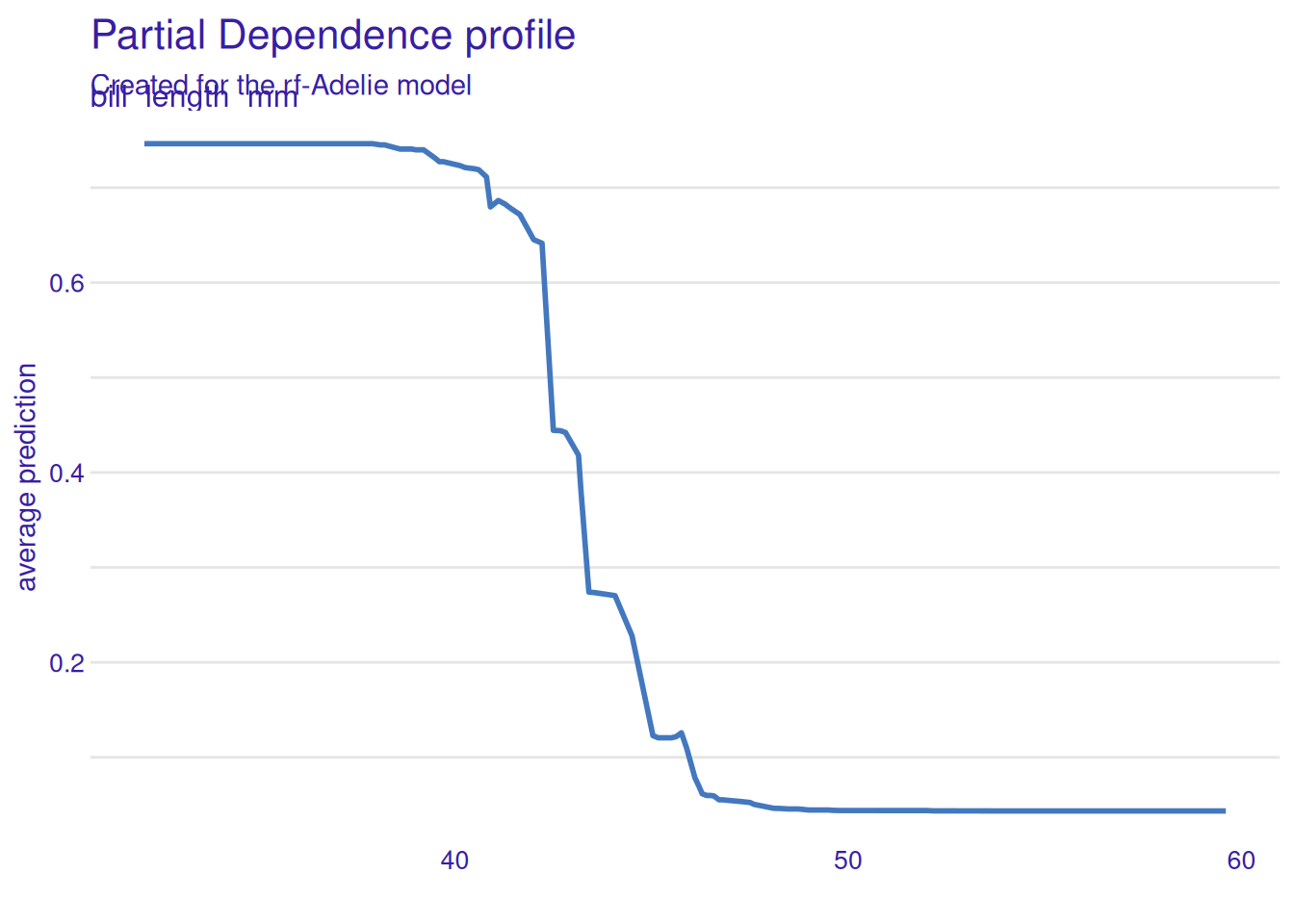
4. Check
A Shapley value for one instance.
pred_fun <- function(model, newdata) predict(model, newdata)$predictions[, "Adelie"]
predictor <- iml::Predictor$new(rf, data = d[, 3:6],
y = as.integer(d$species == "Adelie"),
predict.function = pred_fun)
sh <- iml::Shapley$new(predictor, x.interest = d[1, 3:6])
plot(sh)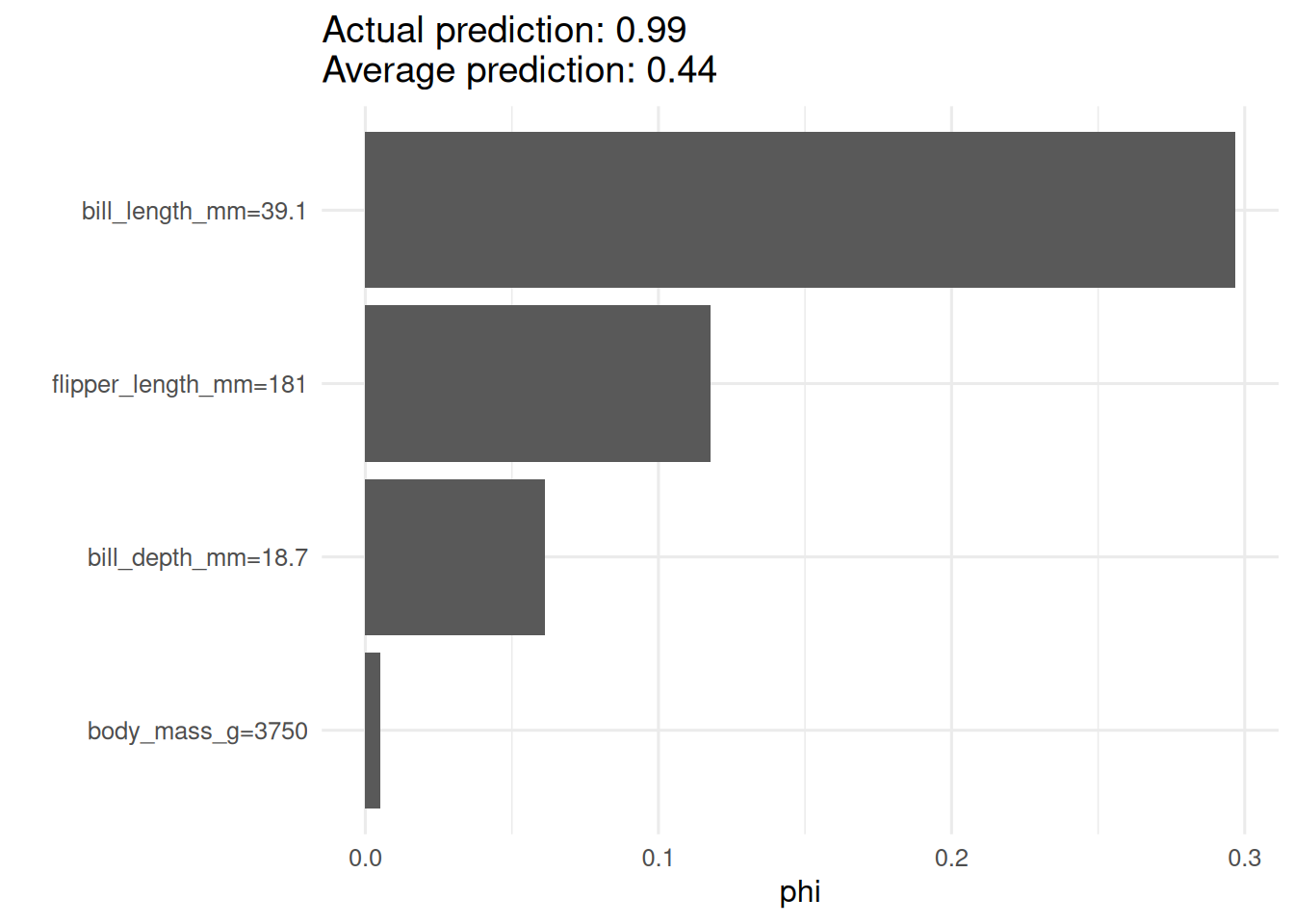
5. Report
On the penguins dataset, a random forest predicting the probability of Adelie species had permutation-importance values dominated by bill length and flipper length. Partial-dependence profiles showed that Adelie probability decreases sharply above a bill length of about 44 mm. For the first instance in the dataset, the Shapley attribution confirmed bill length as the primary contributor.
Report all three artefacts together, and note the correlation structure among the features when you do.
Explain briefly that SHAP values are additive by construction: the sum of contributions plus the baseline equals the prediction.
Common pitfalls
- Treating SHAP values as causal effects.
- Running permutation importance on correlated features without a conditional variant.
- Presenting partial-dependence plots outside the data’s support.
Further reading
- Lundberg SM, Lee S-I (2017), A unified approach to interpreting model predictions.
- Molnar C, Interpretable Machine Learning (online book).
Session info
sessionInfo()R version 4.4.1 (2024-06-14)
Platform: x86_64-pc-linux-gnu
Running under: Ubuntu 24.04.4 LTS
Matrix products: default
BLAS: /usr/lib/x86_64-linux-gnu/openblas-pthread/libblas.so.3
LAPACK: /usr/lib/x86_64-linux-gnu/openblas-pthread/libopenblasp-r0.3.26.so; LAPACK version 3.12.0
locale:
[1] LC_CTYPE=C.UTF-8 LC_NUMERIC=C LC_TIME=C.UTF-8
[4] LC_COLLATE=C.UTF-8 LC_MONETARY=C.UTF-8 LC_MESSAGES=C.UTF-8
[7] LC_PAPER=C.UTF-8 LC_NAME=C LC_ADDRESS=C
[10] LC_TELEPHONE=C LC_MEASUREMENT=C.UTF-8 LC_IDENTIFICATION=C
time zone: UTC
tzcode source: system (glibc)
attached base packages:
[1] stats graphics grDevices utils datasets methods base
other attached packages:
[1] iml_0.11.4 DALEX_2.5.3 ranger_0.18.0 lubridate_1.9.5
[5] forcats_1.0.1 stringr_1.6.0 dplyr_1.2.1 purrr_1.2.2
[9] readr_2.2.0 tidyr_1.3.2 tibble_3.3.1 ggplot2_4.0.3
[13] tidyverse_2.0.0
loaded via a namespace (and not attached):
[1] future_1.70.0 generics_0.1.4 stringi_1.8.7
[4] lattice_0.22-6 listenv_0.10.1 hms_1.1.4
[7] digest_0.6.39 magrittr_2.0.5 evaluate_1.0.5
[10] grid_4.4.1 timechange_0.4.0 RColorBrewer_1.1-3
[13] fastmap_1.2.0 jsonlite_2.0.0 Matrix_1.7-0
[16] backports_1.5.1 scales_1.4.0 palmerpenguins_0.1.1
[19] codetools_0.2-20 cli_3.6.6 rlang_1.2.0
[22] parallelly_1.47.0 withr_3.0.2 yaml_2.3.12
[25] otel_0.2.0 tools_4.4.1 parallel_4.4.1
[28] tzdb_0.5.0 Metrics_0.1.4 checkmate_2.3.4
[31] ingredients_2.3.0 globals_0.19.1 vctrs_0.7.3
[34] R6_2.6.1 lifecycle_1.0.5 htmlwidgets_1.6.4
[37] pkgconfig_2.0.3 pillar_1.11.1 gtable_0.3.6
[40] data.table_1.18.2.1 glue_1.8.1 Rcpp_1.1.1-1.1
[43] xfun_0.57 tidyselect_1.2.1 knitr_1.51
[46] farver_2.1.2 htmltools_0.5.9 labeling_0.4.3
[49] rmarkdown_2.31 compiler_4.4.1 S7_0.2.2