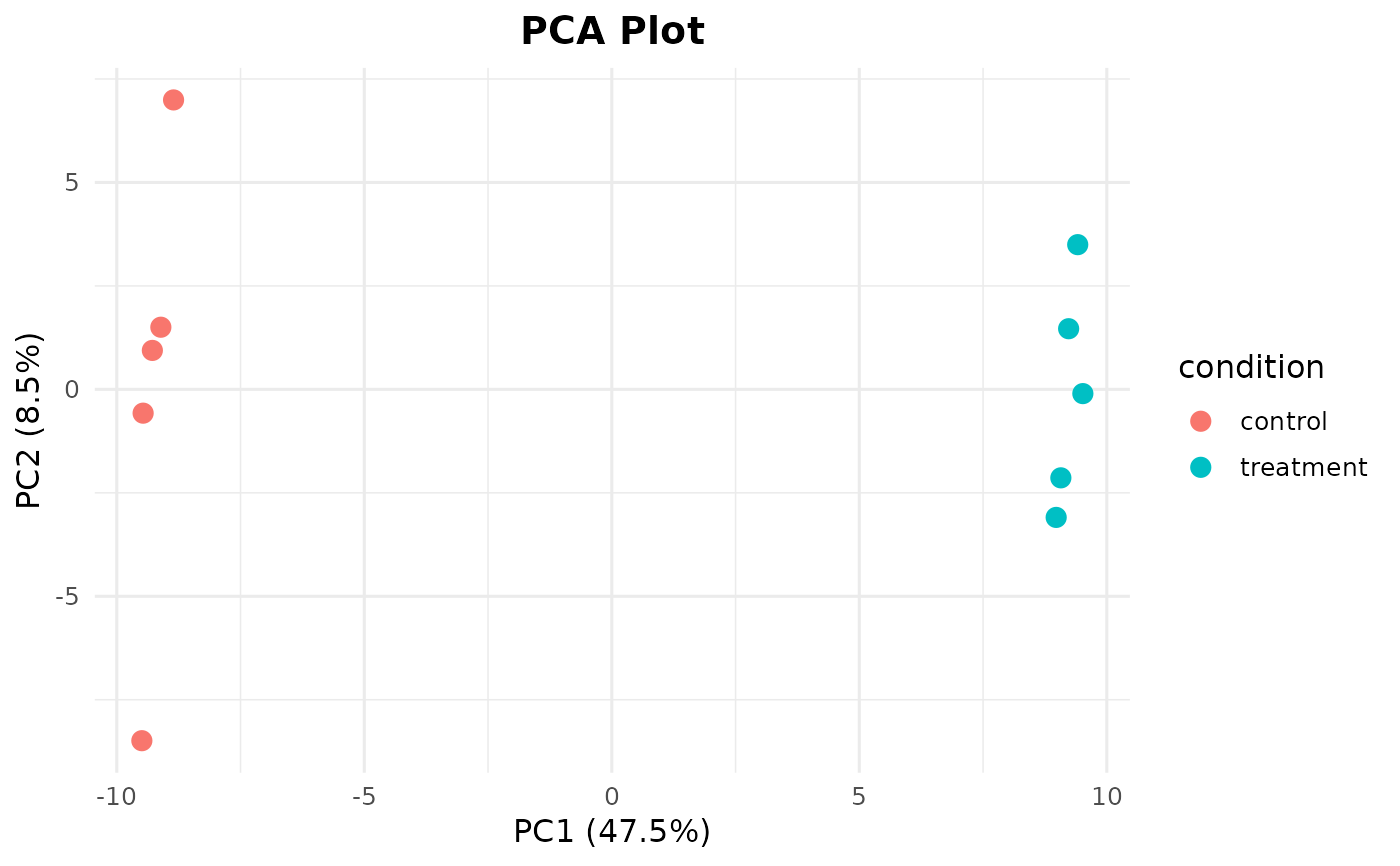
Loads a synthetic RNA-seq count matrix (200 genes x 10 samples) with sample metadata. The first 20 genes have simulated differential expression between control and treatment groups.
Value
A list with components:
- counts
A 200 x 10 integer matrix of raw counts. Rownames are gene symbols, colnames are sample IDs.
- metadata
A data.frame with columns
condition,batch,sex,age. Rownames match colnames ofcounts.
Examples
ex <- bb_example_counts()
dim(ex$counts)
#> [1] 200 10
head(ex$metadata)
#> condition batch sex age
#> Sample_01 control A M 45
#> Sample_02 control B F 52
#> Sample_03 control A M 38
#> Sample_04 control B F 61
#> Sample_05 control A M 55
#> Sample_06 treatment B F 47
# Quick pipeline from example data
cpm <- bb_normalize(ex$counts, method = "cpm")
bb_pca(cpm, ex$metadata, color_by = "condition")