library(tidyverse)
library(palmerpenguins)
library(gtsummary)
set.seed(42)
theme_set(theme_minimal(base_size = 12))Week 2, Session 1 — Descriptive statistics and Table 1
Course 1 — #courses
Workflow labs use the variant template: Goal → Approach → Execution → Check → Report.
Learning objectives
- Match the right summary statistic to the right variable type — means for symmetric continuous, medians for skewed, counts and percentages for categorical.
- Build a publication-ready Table 1 with
gtsummary::tbl_summary(). - Defend the choice of summary with a quick visual check.
Prerequisites
Labs 1.3 through 1.5.
Background
Table 1 is the first real table of almost every clinical paper. It describes the sample: who was in the study, in how many groups, with what characteristics, and — if the study has arms — how balanced those arms are at baseline. A Table 1 that is built thoughtlessly hides an unbalanced comparison; a Table 1 that is built well anchors the rest of the paper.
The right summary depends on the variable. A continuous variable that is roughly symmetric is summarised by its mean and standard deviation; one that is skewed or has heavy tails is summarised by its median and interquartile range. A categorical variable is summarised by counts and percentages. Reporting the median of a balanced continuous variable is not wrong, but it is unusual; reporting the mean of a skewed variable is often misleading.
gtsummary packages these choices into a declarative API. You say which variables to include and which grouping variable to stratify by, and the table is built with sensible defaults. When you need non-standard behaviour, you override per-variable.
Setup
1. Goal
Produce a Table 1 for the penguins dataset stratified by species, and back the choice of summary with a diagnostic plot.
2. Approach
dat <- penguins |>
drop_na(species, sex, bill_length_mm, bill_depth_mm,
flipper_length_mm, body_mass_g)3. Execution
Pick the right summary per variable. First, eyeball the shapes.
dat |>
pivot_longer(c(bill_length_mm, bill_depth_mm,
flipper_length_mm, body_mass_g),
names_to = "variable", values_to = "value") |>
ggplot(aes(value, fill = species)) +
geom_histogram(bins = 20, alpha = 0.6, colour = "grey30",
position = "identity") +
facet_wrap(~ variable, scales = "free") +
labs(x = NULL, y = "Count")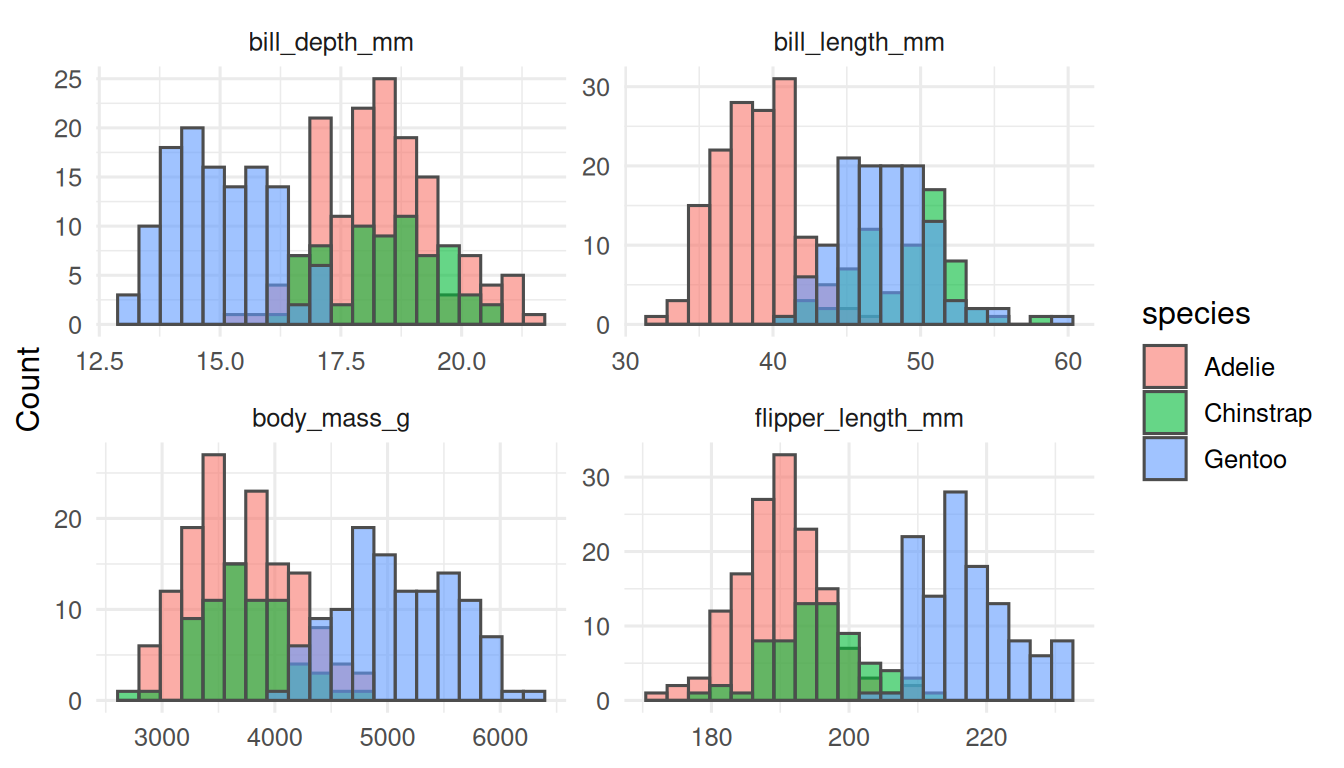
All four continuous variables are approximately symmetric within species, so mean (SD) is defensible. If a variable were skewed, we would switch to median (IQR).
t1 <- dat |>
select(species, sex, bill_length_mm, bill_depth_mm,
flipper_length_mm, body_mass_g) |>
tbl_summary(
by = species,
statistic = list(
all_continuous() ~ "{mean} ({sd})",
all_categorical() ~ "{n} ({p}%)"
),
digits = all_continuous() ~ 1,
label = list(
bill_length_mm ~ "Bill length (mm)",
bill_depth_mm ~ "Bill depth (mm)",
flipper_length_mm ~ "Flipper length (mm)",
body_mass_g ~ "Body mass (g)",
sex ~ "Sex"
)
) |>
add_overall() |>
add_p()
t1| Characteristic | Overall N = 3331 |
Adelie N = 1461 |
Chinstrap N = 681 |
Gentoo N = 1191 |
p-value2 |
|---|---|---|---|---|---|
| Sex | >0.9 | ||||
| female | 165 (50%) | 73 (50%) | 34 (50%) | 58 (49%) | |
| male | 168 (50%) | 73 (50%) | 34 (50%) | 61 (51%) | |
| Bill length (mm) | 44.0 (5.5) | 38.8 (2.7) | 48.8 (3.3) | 47.6 (3.1) | <0.001 |
| Bill depth (mm) | 17.2 (2.0) | 18.3 (1.2) | 18.4 (1.1) | 15.0 (1.0) | <0.001 |
| Flipper length (mm) | 201.0 (14.0) | 190.1 (6.5) | 195.8 (7.1) | 217.2 (6.6) | <0.001 |
| Body mass (g) | 4,207.1 (805.2) | 3,706.2 (458.6) | 3,733.1 (384.3) | 5,092.4 (501.5) | <0.001 |
| 1 n (%); Mean (SD) | |||||
| 2 Pearson’s Chi-squared test; Kruskal-Wallis rank sum test | |||||
add_p() will run a t-test or ANOVA on continuous variables and a chi-square or Fisher’s exact test on categorical, with sensible defaults for sample size.
4. Check
Validate the gtsummary output against a hand computation.
dat |>
group_by(species) |>
summarise(
n = n(),
mean_mass = mean(body_mass_g),
sd_mass = sd(body_mass_g),
.groups = "drop"
)# A tibble: 3 × 4
species n mean_mass sd_mass
<fct> <int> <dbl> <dbl>
1 Adelie 146 3706. 459.
2 Chinstrap 68 3733. 384.
3 Gentoo 119 5092. 501.The group means and standard deviations must match the numbers in the body-mass row of the Table 1.
5. Report
Table 1 describes 333 penguins from three species in the Palmer Archipelago. Body mass was highest in Gentoo (5092 ± 501 g) and lower in Adelie and Chinstrap. All four continuous variables were approximately symmetric within species and are summarised as mean (SD); sex was approximately balanced within each species.
Table 1 is not a tool for hypothesis testing. The p-values in the rightmost column describe the data, not the biology. A statistically-significant difference between arms at baseline in a randomised trial is, by construction, a chance finding.
Stress the distinction between Table 1 as description and Table 2 as inference. The audience will carry the habit into their own writing.
Common pitfalls
- Reporting a mean for a skewed variable.
- Using Table 1 as a place to “prove balance” with p-values in a randomised trial.
- Suppressing the missing-data row. Always report it.
- Forgetting that
tbl_summarydrops NAs by default; check the count at the bottom of each column.
Further reading
- Sjoberg DD et al. Reproducible summary tables with the gtsummary package. R Journal.
- Altman DG. Practical Statistics for Medical Research, chapter on describing data.
Session info
sessionInfo()R version 4.4.1 (2024-06-14)
Platform: x86_64-pc-linux-gnu
Running under: Ubuntu 24.04.4 LTS
Matrix products: default
BLAS: /usr/lib/x86_64-linux-gnu/openblas-pthread/libblas.so.3
LAPACK: /usr/lib/x86_64-linux-gnu/openblas-pthread/libopenblasp-r0.3.26.so; LAPACK version 3.12.0
locale:
[1] LC_CTYPE=C.UTF-8 LC_NUMERIC=C LC_TIME=C.UTF-8
[4] LC_COLLATE=C.UTF-8 LC_MONETARY=C.UTF-8 LC_MESSAGES=C.UTF-8
[7] LC_PAPER=C.UTF-8 LC_NAME=C LC_ADDRESS=C
[10] LC_TELEPHONE=C LC_MEASUREMENT=C.UTF-8 LC_IDENTIFICATION=C
time zone: UTC
tzcode source: system (glibc)
attached base packages:
[1] stats graphics grDevices utils datasets methods base
other attached packages:
[1] gtsummary_2.5.0 palmerpenguins_0.1.1 lubridate_1.9.5
[4] forcats_1.0.1 stringr_1.6.0 dplyr_1.2.1
[7] purrr_1.2.2 readr_2.2.0 tidyr_1.3.2
[10] tibble_3.3.1 ggplot2_4.0.3 tidyverse_2.0.0
loaded via a namespace (and not attached):
[1] gt_1.3.0 utf8_1.2.6 sass_0.4.10 generics_0.1.4
[5] xml2_1.5.2 stringi_1.8.7 hms_1.1.4 digest_0.6.39
[9] magrittr_2.0.5 evaluate_1.0.5 grid_4.4.1 timechange_0.4.0
[13] RColorBrewer_1.1-3 cards_0.7.1 fastmap_1.2.0 jsonlite_2.0.0
[17] cardx_0.3.2 backports_1.5.1 scales_1.4.0 cli_3.6.6
[21] rlang_1.2.0 litedown_0.9 commonmark_2.0.0 base64enc_0.1-6
[25] withr_3.0.2 yaml_2.3.12 otel_0.2.0 tools_4.4.1
[29] tzdb_0.5.0 broom_1.0.12 vctrs_0.7.3 R6_2.6.1
[33] lifecycle_1.0.5 fs_2.1.0 htmlwidgets_1.6.4 pkgconfig_2.0.3
[37] pillar_1.11.1 gtable_0.3.6 glue_1.8.1 xfun_0.57
[41] tidyselect_1.2.1 knitr_1.51 farver_2.1.2 htmltools_0.5.9
[45] rmarkdown_2.31 labeling_0.4.3 compiler_4.4.1 S7_0.2.2
[49] markdown_2.0