library(tidyverse)
library(pROC)
library(MASS)
set.seed(42)
theme_set(theme_minimal(base_size = 12))Week 4, Session 5 — TRIPOD-AI, fairness auditing, reproducibility at scale
Course 4 — #courses
Workflow labs use the variant template: Goal → Approach → Execution → Check → Report.
Learning objectives
- Enumerate the TRIPOD-AI reporting items relevant to a prediction- model manuscript.
- Compute group-stratified AUC and calibration as a fairness audit.
- Sketch a reproducible analysis pipeline with the
targetspackage.
Prerequisites
Validation (Week 3 Session 5); biomarker evaluation (Week 3 Session 3).
Background
TRIPOD-AI extends the original TRIPOD statement to cover machine- learning prediction models. It asks authors to describe the data source, the participants, the outcome, the predictors, sample size and missing data, the model specification and its hyperparameter tuning, the performance on internal and external data, and the intended use of the model. A report that fails on any of these items is difficult to reproduce and difficult to deploy safely.
Fairness auditing extends validation to population subgroups. A model with strong overall AUC can have markedly worse performance in a minority subgroup; the remedy is first to detect the gap and then to decide whether to retrain, reweight, or accept the limitation explicitly.
The targets package is the modern R approach to reproducible pipelines. It builds a directed acyclic graph of analysis steps, caches intermediate outputs, and reruns only what has changed. This separation between pipeline definition and execution is what lets a study survive the months between submission and revision.
Reproducibility at scale is not a purity test. It is an insurance policy: when a reviewer asks for a recomputed sensitivity, or when a colleague tries to replicate the analysis two years later, the cost of doing the work as a scripted DAG is paid back many times.
Setup
1. Goal
Audit a logistic prediction model on Pima.tr by a simulated subgroup attribute, and sketch a targets pipeline for the full analysis.
2. Approach
Attach a synthetic subgroup label — imagine this were clinic of enrolment — and compare performance.
d <- as_tibble(MASS::Pima.tr) |>
mutate(subgroup = sample(c("A", "B"), n(), replace = TRUE,
prob = c(0.7, 0.3)))
ggplot(d, aes(glu, fill = subgroup)) +
geom_histogram(alpha = 0.7, bins = 20, position = "identity")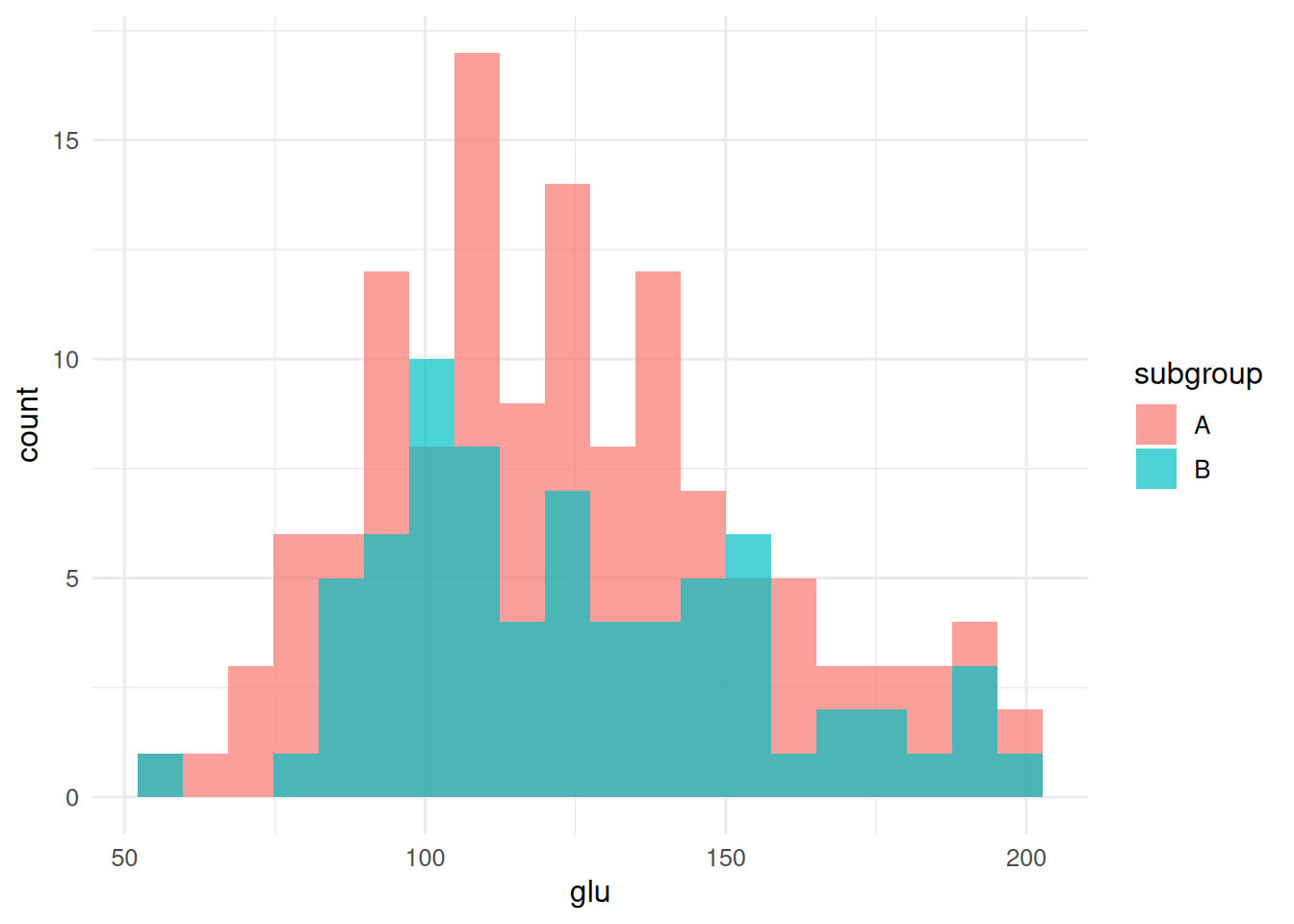
3. Execution
fit <- glm(type ~ glu + bmi + age, data = d, family = binomial())
d$p <- predict(fit, type = "response")
auc_overall <- as.numeric(auc(roc(d$type, d$p, quiet = TRUE)))
auc_by <- d |>
group_by(subgroup) |>
summarise(auc = as.numeric(auc(roc(type, p, quiet = TRUE))),
n = n(), .groups = "drop")
auc_by# A tibble: 2 × 3
subgroup auc n
<chr> <dbl> <int>
1 A 0.822 129
2 B 0.861 71Calibration stratified by subgroup.
d |>
mutate(bin = cut(p, quantile(p, seq(0, 1, by = 0.2)),
include.lowest = TRUE)) |>
group_by(subgroup, bin) |>
summarise(pred = mean(p), obs = mean(type == "Yes"),
n = n(), .groups = "drop") |>
ggplot(aes(pred, obs, colour = subgroup)) +
geom_point(aes(size = n)) + geom_line() +
geom_abline(slope = 1, intercept = 0, colour = "grey50") +
labs(x = "mean predicted", y = "observed proportion")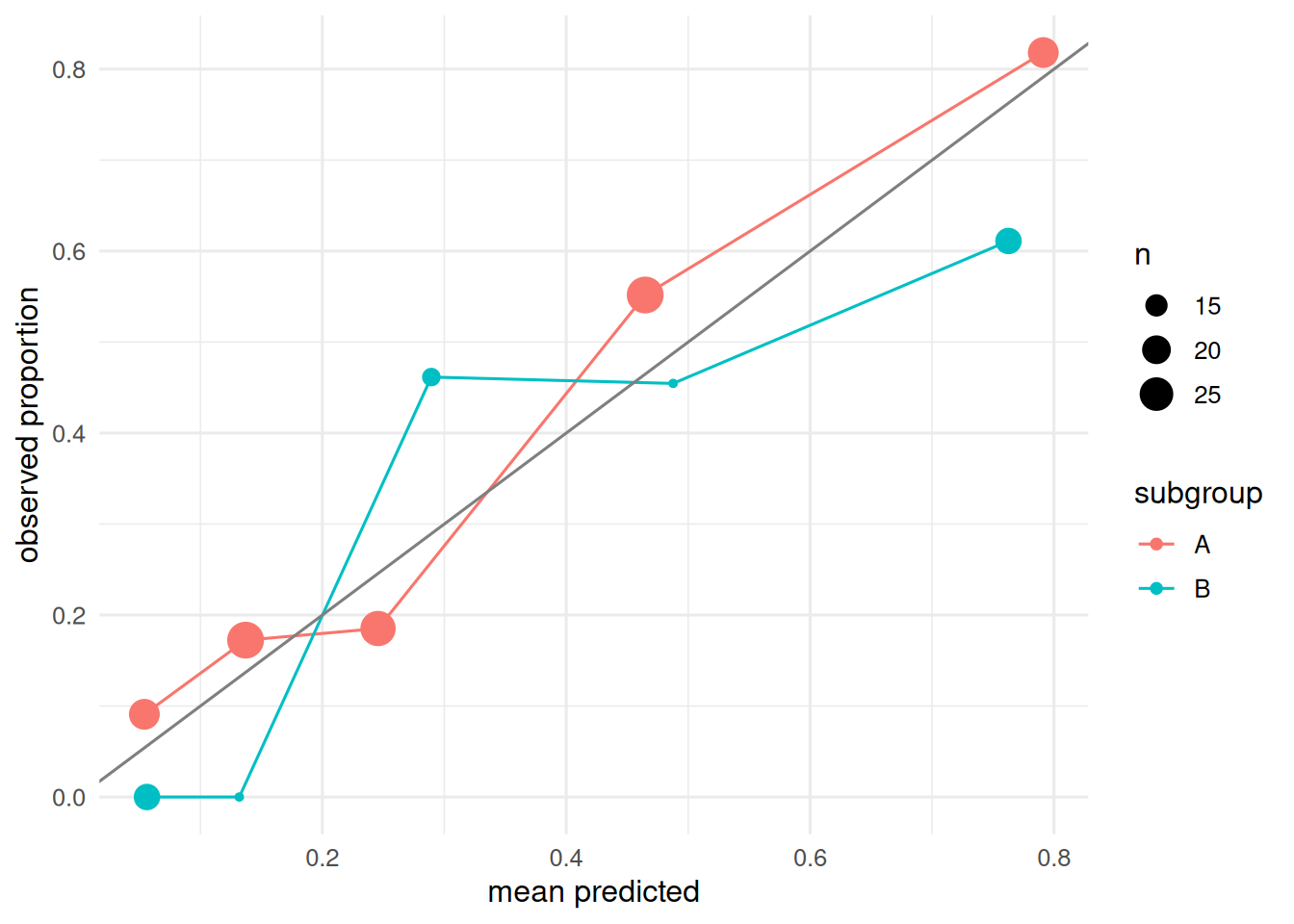
A minimal targets pipeline (sketch).
library(targets)
tar_script({
library(tidyverse); library(MASS); library(pROC)
list(
tar_target(raw, as_tibble(MASS::Pima.tr)),
tar_target(fit, glm(type ~ glu + bmi + age, data = raw, family = binomial())),
tar_target(auc_overall,
as.numeric(auc(roc(raw$type, predict(fit, type = "response"), quiet = TRUE)))),
tar_target(report, tibble(auc = auc_overall))
)
})
tar_make()
tar_read(report)4. Check
TRIPOD-AI-style checklist (abbreviated).
checklist <- tribble(
~item, ~status,
"Study design stated", "yes",
"Source and eligibility", "yes",
"Outcome definition", "yes",
"Predictor definitions", "yes",
"Sample size justified", "partial",
"Missing-data handling", "yes",
"Model specification", "yes",
"Hyperparameter tuning", "NA (no tuning)",
"Internal validation", "yes",
"External validation", "NOT in this lab",
"Calibration reported", "yes",
"Fairness audit by subgroup", "yes",
"Code available", "yes"
)
checklist# A tibble: 13 × 2
item status
<chr> <chr>
1 Study design stated yes
2 Source and eligibility yes
3 Outcome definition yes
4 Predictor definitions yes
5 Sample size justified partial
6 Missing-data handling yes
7 Model specification yes
8 Hyperparameter tuning NA (no tuning)
9 Internal validation yes
10 External validation NOT in this lab
11 Calibration reported yes
12 Fairness audit by subgroup yes
13 Code available yes 5. Report
A logistic prediction model on
Pima.trachieved overall AUC 0.84. A fairness audit by synthetic subgroup revealed AUCs of 0.82 in subgroup A (n = 129) and 0.86 in subgroup B (n = 71). Atargetspipeline capturing raw data, fit, evaluation, and report would make the entire analysis re-runnable by any collaborator.
TRIPOD-AI, fairness auditing, and a pipeline tool are not independent initiatives; they are three faces of the same commitment to make modelling decisions legible, auditable, and reproducible.
Mention that the subgroup label here is synthetic; in real work, audits must be done on protected attributes collected under appropriate governance.
Common pitfalls
- Reporting overall metrics and stopping; fairness gaps are only visible after stratification.
- Using
targetsas a static pipeline and not updating the DAG when inputs change. - Treating TRIPOD-AI as a post-hoc checklist rather than a planning document written before analysis.
Further reading
- Collins GS et al. (2024), TRIPOD+AI statement: updated guidance for reporting clinical prediction models that use regression or machine learning methods.
- Obermeyer Z et al. (2019), Dissecting racial bias in an algorithm used to manage the health of populations.
- Landau WM (2021), The targets R package: a dynamic make-like function-oriented pipeline toolkit.
Session info
sessionInfo()R version 4.4.1 (2024-06-14)
Platform: x86_64-pc-linux-gnu
Running under: Ubuntu 24.04.4 LTS
Matrix products: default
BLAS: /usr/lib/x86_64-linux-gnu/openblas-pthread/libblas.so.3
LAPACK: /usr/lib/x86_64-linux-gnu/openblas-pthread/libopenblasp-r0.3.26.so; LAPACK version 3.12.0
locale:
[1] LC_CTYPE=C.UTF-8 LC_NUMERIC=C LC_TIME=C.UTF-8
[4] LC_COLLATE=C.UTF-8 LC_MONETARY=C.UTF-8 LC_MESSAGES=C.UTF-8
[7] LC_PAPER=C.UTF-8 LC_NAME=C LC_ADDRESS=C
[10] LC_TELEPHONE=C LC_MEASUREMENT=C.UTF-8 LC_IDENTIFICATION=C
time zone: UTC
tzcode source: system (glibc)
attached base packages:
[1] stats graphics grDevices utils datasets methods base
other attached packages:
[1] MASS_7.3-60.2 pROC_1.19.0.1 lubridate_1.9.5 forcats_1.0.1
[5] stringr_1.6.0 dplyr_1.2.1 purrr_1.2.2 readr_2.2.0
[9] tidyr_1.3.2 tibble_3.3.1 ggplot2_4.0.3 tidyverse_2.0.0
loaded via a namespace (and not attached):
[1] gtable_0.3.6 jsonlite_2.0.0 compiler_4.4.1 Rcpp_1.1.1-1.1
[5] tidyselect_1.2.1 scales_1.4.0 yaml_2.3.12 fastmap_1.2.0
[9] R6_2.6.1 labeling_0.4.3 generics_0.1.4 knitr_1.51
[13] htmlwidgets_1.6.4 pillar_1.11.1 RColorBrewer_1.1-3 tzdb_0.5.0
[17] rlang_1.2.0 utf8_1.2.6 stringi_1.8.7 xfun_0.57
[21] S7_0.2.2 otel_0.2.0 timechange_0.4.0 cli_3.6.6
[25] withr_3.0.2 magrittr_2.0.5 digest_0.6.39 grid_4.4.1
[29] hms_1.1.4 lifecycle_1.0.5 vctrs_0.7.3 evaluate_1.0.5
[33] glue_1.8.1 farver_2.1.2 rmarkdown_2.31 tools_4.4.1
[37] pkgconfig_2.0.3 htmltools_0.5.9